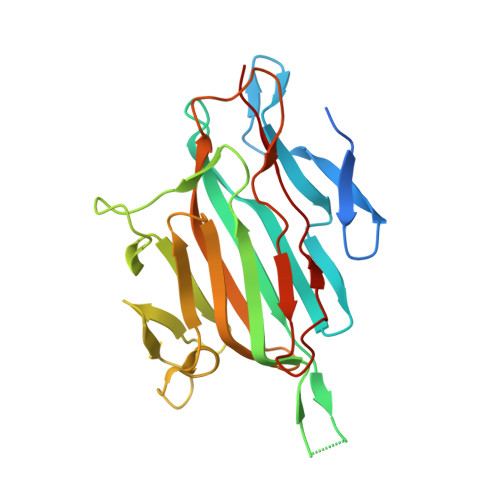
Autocatalytic association of proteins by covalent bond formation: a Bio Molecular Welding toolbox derived from a bacterial adhesin.
Bonnet, J., Cartannaz, J., Tourcier, G., Contreras-Martel, C., Kleman, J.P., Morlot, C., Vernet, T., Di Guilmi, A.M.(2017) Sci Rep 7: 43564-43564
- PubMed: 28252635 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep43564
- Primary Citation Related Structures:
5MKC - PubMed Abstract:
Unusual intramolecular cross-links present in adhesins from Gram-positive bacteria have been used to develop a generic process amenable to biotechnology applications. Based on the crystal structure of RrgA, the Streptococcus pneumoniae pilus adhesin, we provide evidence that two engineered protein fragments retain their ability to associate covalently with high specificity, in vivo and in vitro, once isolated from the parent protein. We determined the optimal conditions for the assembly of the complex and we solved its crystal structure at 2 Å. Furthermore, we demonstrate biotechnological applications related to antibody production, nanoassembly and cell-surface labeling based on this process we named Bio Molecular Welding.
- Institut de Biologie Structurale (IBS), Univ. Grenoble Alpes, CEA, CNRS, 38044 Grenoble, France.
Organizational Affiliation: