Thrombin Mutant A190S in complex with (S) -1 - ((R) -2-amino-3,3-diphenylpropanoyl) -N- (3-chlorobenzyl) pyrrolidine-2-carboxamide
Marca, A., Ngo, K., Sandner, A., Heine, A., Klebe, G.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Thrombin light chain | A [auth L] | 36 | Homo sapiens | Mutation(s): 0 Gene Names: F2 EC: 3.4.21.5 |  |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P00734 (Homo sapiens) Explore P00734 Go to UniProtKB: P00734 | |||||
PHAROS: P00734 GTEx: ENSG00000180210 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00734 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Thrombin heavy chain | B [auth H] | 259 | Homo sapiens | Mutation(s): 0 Gene Names: F2 EC: 3.4.21.5 | 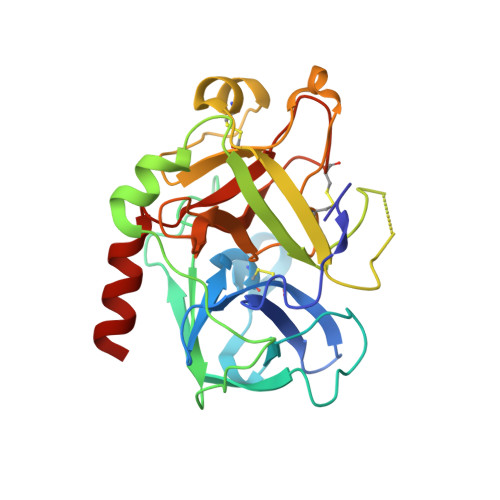 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P00734 (Homo sapiens) Explore P00734 Go to UniProtKB: P00734 | |||||
PHAROS: P00734 GTEx: ENSG00000180210 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00734 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hirudin variant-2 | C [auth D] | 12 | Hirudo medicinalis | Mutation(s): 0 |  |
UniProt | |||||
Find proteins for P09945 (Hirudo medicinalis) Explore P09945 Go to UniProtKB: P09945 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P09945 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 23U Query on 23U | D [auth H] | beta-phenyl-D-phenylalanyl-N-(3-chlorobenzyl)-L-prolinamide C27 H28 Cl N3 O2 LJTKFVOSAVCILF-UKILVPOCSA-N |  | ||
| PO4 Query on PO4 | F [auth H] | PHOSPHATE ION O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-K |  | ||
| GOL Query on GOL | G [auth H], H, I [auth H], J [auth H], M [auth H] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| DMS Query on DMS | E [auth H] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| NA Query on NA | K [auth H], L [auth H] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| TYS Query on TYS | C [auth D] | L-PEPTIDE LINKING | C9 H11 N O6 S |  | TYR |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 69.492 | α = 90 |
| b = 71.613 | β = 99.73 |
| c = 72.129 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| Coot | model building |
| XDS | data scaling |
| XDS | data reduction |
| PHASER | phasing |