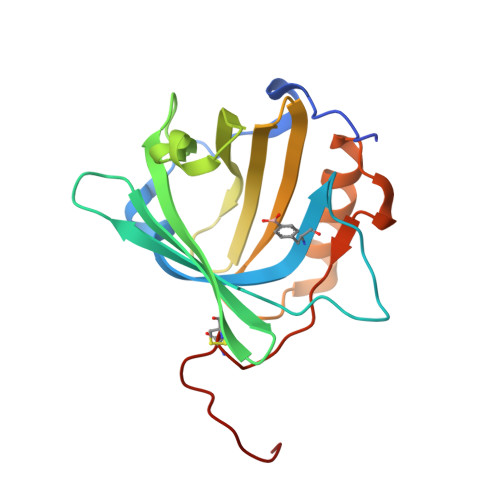
Rational Design of an Anticalin-Type Sugar-Binding Protein Using a Genetically Encoded Boronate Side Chain.
Edwardraja, S., Eichinger, A., Theobald, I., Sommer, C.A., Reichert, A.J., Skerra, A.(2017) ACS Synth Biol 6: 2241-2247
- PubMed: 28937743 Search on PubMed
- DOI: https://doi.org/10.1021/acssynbio.7b00199
- Primary Citation Related Structures:
5MHH - PubMed Abstract:
The molecular recognition of carbohydrates plays a fundamental role in many biological processes. However, the development of carbohydrate-binding reagents for biomedical research and use poses a challenge due to the generally poor affinity of proteins toward sugars in aqueous solution. Here, we describe the effective molecular recognition of pyranose monosaccharides (in particular, galactose and mannose) by a rationally designed protein receptor based on the human lipocalin scaffold (Anticalin). Complexation relies on reversible covalent cis-diol boronate diester formation with a genetically encoded l-boronophenylalanine (Bpa) residue which was incorporated as a non-natural amino acid at a sterically permissive position in the ligand pocket of the Anticalin, as confirmed by X-ray crystallography. Compared with the metal-ion and/or avidity-dependent oligovalent lectins that prevail in nature, our approach offers a novel and promising route to generate tight sugar-binding reagents both as research reagents and for biomedical applications.
- Munich Center for Integrated Protein Science (CIPS-M) and Lehrstuhl für Biologische Chemie, Technische Universität München , Emil-Erlenmeyer-Forum 5, 85354 Freising (Weihenstephan), Germany.
Organizational Affiliation: