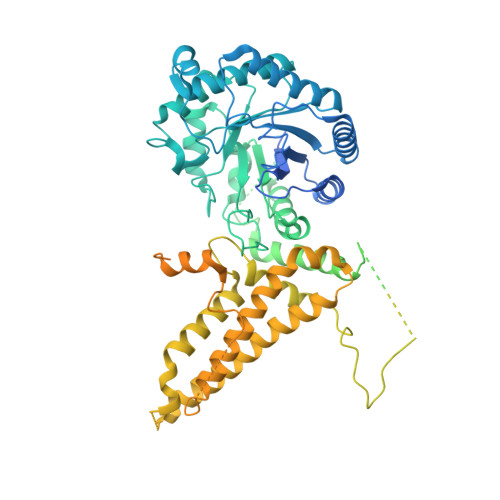
Structural and functional insight into human O-GlcNAcase.
Roth, C., Chan, S., Offen, W.A., Hemsworth, G.R., Willems, L.I., King, D.T., Varghese, V., Britton, R., Vocadlo, D.J., Davies, G.J.(2017) Nat Chem Biol 13: 610-612
- PubMed: 28346405 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.2358
- Primary Citation Related Structures:
5M7R, 5M7S, 5M7T, 5M7U - PubMed Abstract:
O-GlcNAc hydrolase (OGA) removes O-linked N-acetylglucosamine (O-GlcNAc) from a myriad of nucleocytoplasmic proteins. Through co-expression and assembly of OGA fragments, we determined the three-dimensional structure of human OGA, revealing an unusual helix-exchanged dimer that lays a structural foundation for an improved understanding of substrate recognition and regulation of OGA. Structures of OGA in complex with a series of inhibitors define a precise blueprint for the design of inhibitors that have clinical value.
- York Structural Biology Laboratory, Department of Chemistry University of York, York, UK.
Organizational Affiliation: