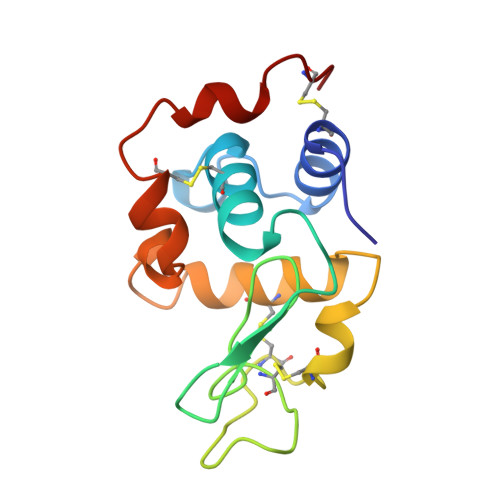
Studies of monoclinic hen egg-white lysozyme. IV. X-ray refinement at 1.8 A resolution and a comparison of the variable regions in the polymorphic forms.
Rao, S.T., Sundaralingam, M.(1996) Acta Crystallogr D Biol Crystallogr 52: 170-175
- PubMed: 15299739 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444995009504
- Primary Citation Related Structures:
5LYM - PubMed Abstract:
Monoclinic crystals of hen egg-white lysozyme (E.C. 3.2.1.17, HEL) grown at low pH in the presence of NaNO(3) belong to space group P2(1) with unit-cell dimensions, a = 28.0, b = 62.5, c = 60.9 A and beta= 90.8 degrees with two molecules in the asymmetric unit. 1.8 A resolution intensity data, collected on a CAD-4 diffractometer, contained 17 524 reflections with F > 3sigma (93% complete). Our earlier preliminary 1.8 A model was refitted and refined using X-PLOR to an R value of 0.189. The deviations in the model from ideal geometry are 0.013 A in bond lengths and 2.8 degrees in bond angles. The r.m.s. deviation in the backbone atoms between the two molecules is 0.42 A. A comparison of HEL in different polymorphic crystal forms reveals that the prominent structural variability among them resides in two exposed regions 45-50 and 65-73 which are also regions of lattice contacts.
- Laboratory of Biological Macromolecular Structure, Department of Chemistry, The Ohio State University, Columbus 43210, USA.
Organizational Affiliation: