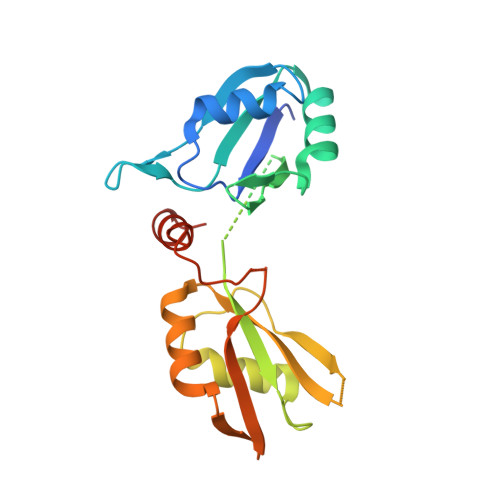
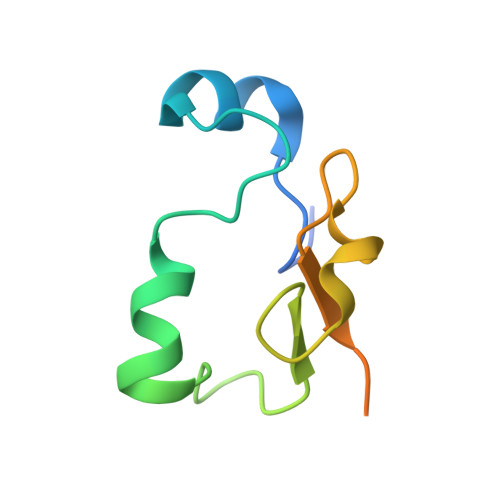
Crystal structure of U2 snRNP SF3b components: Hsh49p in complex with Cus1p-binding domain.
van Roon, A.M., Oubridge, C., Obayashi, E., Sposito, B., Newman, A.J., Seraphin, B., Nagai, K.(2017) RNA 23: 968-981
- PubMed: 28348170 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.059378.116
- Primary Citation Related Structures:
5LSB, 5LSL - PubMed Abstract:
Spliceosomal proteins Hsh49p and Cus1p are components of SF3b, which together with SF3a, Msl1p/Lea1p, Sm proteins, and U2 snRNA, form U2 snRNP, which plays a crucial role in pre-mRNA splicing. Hsh49p, comprising two RRMs, forms a heterodimer with Cus1p. We determined the crystal structures of Saccharomyces cerevisiae full-length Hsh49p as well as its RRM1 in complex with a minimal binding region of Cus1p (residues 290-368). The structures show that the Cus1 fragment binds to the α-helical surface of Hsh49p RRM1, opposite the four-stranded β-sheet, leaving the canonical RNA-binding surface available to bind RNA. Hsh49p binds the 5' end region of U2 snRNA via RRM1. Its affinity is increased in complex with Cus1(290-368)p, partly because an extended RNA-binding surface forms across the protein-protein interface. The Hsh49p RRM1-Cus1(290-368)p structure fits well into cryo-EM density of the B act spliceosome, corroborating the biological relevance of our crystal structure.
- MRC Laboratory of Molecular Biology, Cambridge CB2 0QH, United Kingdom.
Organizational Affiliation: