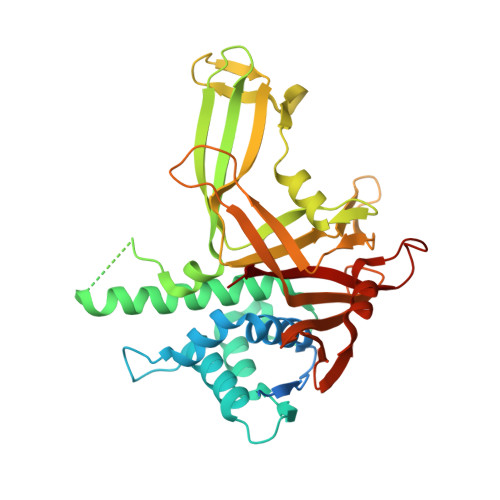
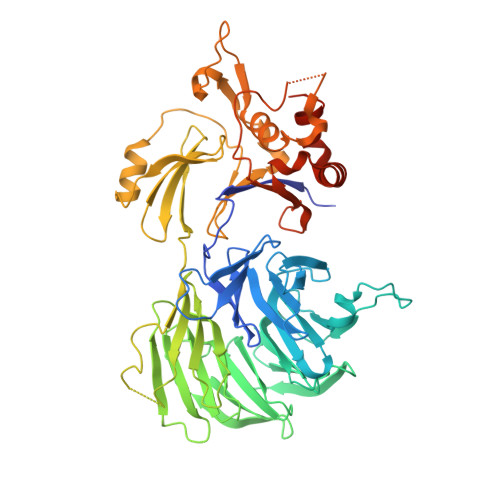
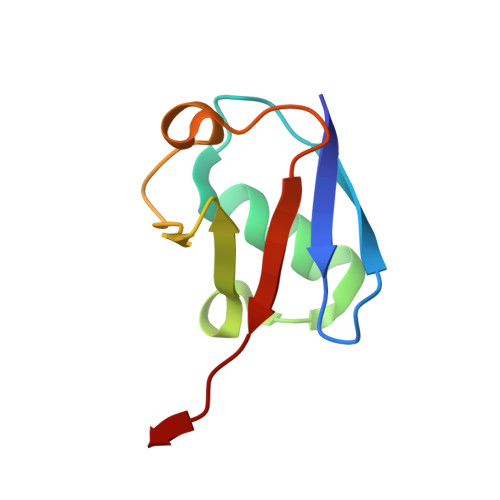
A conserved two-step binding for the UAF1 regulator to the USP12 deubiquitinating enzyme.
Dharadhar, S., Clerici, M., van Dijk, W.J., Fish, A., Sixma, T.K.(2016) J Struct Biol 196: 437-447
- PubMed: 27650958 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2016.09.011
- Primary Citation Related Structures:
5L8E, 5L8H, 5L8W - PubMed Abstract:
Regulation of deubiquitinating enzyme (DUB) activity is an essential step for proper function of cellular ubiquitin signals. UAF1 is a WD40 repeat protein, which binds and activates three important DUBs, USP1, USP12 and USP46. Here, we report the crystal structure of the USP12-Ub/UAF1 complex at a resolution of 2.8Å and of UAF1 at 2.3Å. In the complex we find two potential sites for UAF1 binding, analogous to what was seen in a USP46/UAF1 complex. In line with these observed dual binding states, we show here that USP12/UAF1 complex has 1:2 stoichiometry in solution, with a two-step binding at 4nM and 325nM respectively. Mutagenesis studies show that the fingers sub-domain of USP12 interacts with UAF1 to form the high affinity interface. Our activation studies confirm that the high affinity binding is important for activation while the second UAF1 binding does not affect activation. Nevertheless, we show that this two step binding is conserved in the well-studied USP12 paralog, USP1. Our results highlight the interfaces essential for regulation of USP12 activity and show a conserved second binding of UAF1 which could be important for regulatory functions independent of USP12 activity.
- Division of Biochemistry and CGC.nl, Netherlands Cancer Institute, Plesmanlaan 121, 1066 CX Amsterdam, The Netherlands.
Organizational Affiliation: