The Leaderless Bacteriocin Enterocin K1 Is Highly Potent against Enterococcus faecium: A Study on Structure, Target Spectrum and Receptor.
Ovchinnikov, K.V., Kristiansen, P.E., Straume, D., Jensen, M.S., Aleksandrzak-Piekarczyk, T., Nes, I.F., Diep, D.B.(2017) Front Microbiol 8: 774-774
- PubMed: 28515717 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fmicb.2017.00774
- Primary Citation Related Structures:
5L82 - PubMed Abstract:
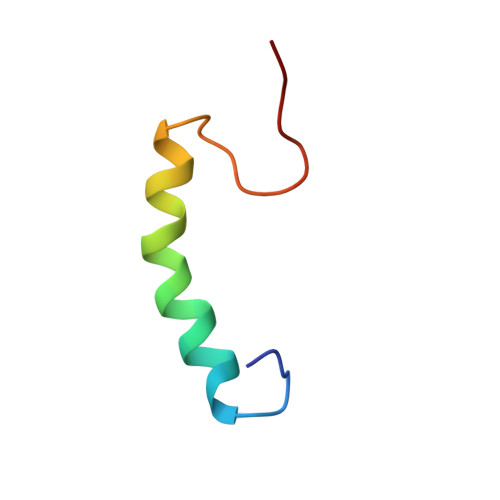
Enterocin K1 (EntK1), enterocin EJ97 (EntEJ97), and LsbB are three sequence related leaderless bacteriocins. Yet LsbB kills only lactococci while EntK1 and EntEJ97 target wider spectra with EntK1 being particularly active against Enterococcus faecium , including nosocomial multidrug resistant isolates. NMR study of EntK1 showed that it had a structure very similar to LsbB - both having an amphiphilic N-terminal α-helix and an unstructured C-terminus. The α-helix in EntK1 is, however, about 3-4 residues longer than that of LsbB. Enterococcal mutants highly resistant to EntEJ97 and EntK1 were found to have mutations within rseP , a gene encoding a stress response membrane-bound Zn-dependent protease. Heterologous expression of the enterococcal rseP rendered resistant cells of Streptococcus pneumoniae sensitive to EntK1 and EntEJ97, suggesting that RseP likely serves as the receptor for EntK1 and EntEJ97. It was also shown that the conserved proteolytic active site in E. faecalis RseP is partly required for EntK1 and EntEJ97 activity, since alanine substitutions of its conserved residues (HExxH) reduced the sensitivity of the clones to the bacteriocins. RseP is known to be involved in bacterial stress response. As expected, the growth of resistant mutants with mutations within rseP was severely affected when they were exposed to higher (stressing) growth temperatures, e.g., at 45°C, at which wild type cells still grew well. These findings allow us to design a hurdle strategy with a combination of the bacteriocin(s) and higher temperature that effectively kills bacteriocin sensitive bacteria and prevents the development of resistant cells.
- Department of Chemistry, Biotechnology and Food Science, Norwegian University of Life SciencesÅs, Norway.
Organizational Affiliation: