Tyrosine sulfation modulates activity of tick-derived thrombin inhibitors.
Thompson, R.E., Liu, X., Ripoll-Rozada, J., Alonso-Garcia, N., Parker, B.L., Pereira, P.J.B., Payne, R.J.(2017) Nat Chem 9: 909-917
- PubMed: 28837178 Search on PubMed
- DOI: https://doi.org/10.1038/nchem.2744
- Primary Citation Related Structures:
5L6N - PubMed Abstract:
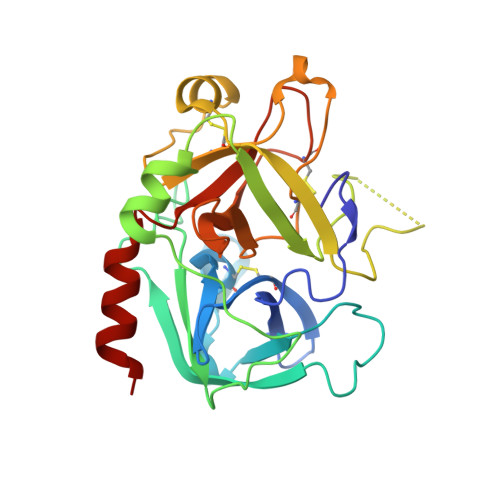
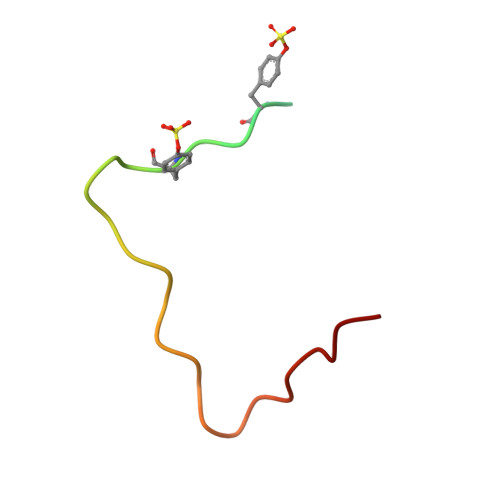
Madanin-1 and chimadanin are two small cysteine-free thrombin inhibitors that facilitate blood feeding in the tick Haemaphysalis longicornis. Here, we report a post-translational modification-tyrosine sulfation-of these two proteins that is critical for potent anti-thrombotic and anticoagulant activity. Inhibitors produced in baculovirus-infected insect cells displayed heterogeneous sulfation of two tyrosine residues within each of the proteins. One-pot ligation-desulfurization chemistry enabled access to homogeneous samples of all possible sulfated variants of the proteins. Tyrosine sulfation of madanin-1 and chimadanin proved crucial for thrombin inhibitory activity, with the doubly sulfated variants three orders of magnitude more potent than the unmodified inhibitors. The three-dimensional structure of madanin-1 in complex with thrombin revealed a unique mode of inhibition, with the sulfated tyrosine residues binding to the basic exosite II of the protease. The importance of tyrosine sulfation within this family of thrombin inhibitors, together with their unique binding mode, paves the way for the development of anti-thrombotic drug leads based on these privileged scaffolds.
- School of Chemistry, The University of Sydney, Sydney, New South Wales 2006, Australia.
Organizational Affiliation: