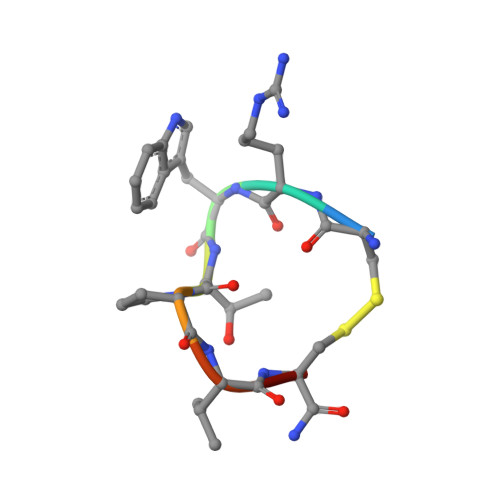
Structure and activity of contryphan-Vc2: Importance of the d-amino acid residue.
Drane, S.B., Robinson, S.D., MacRaild, C.A., Chhabra, S., Chittoor, B., Morales, R.A., Leung, E.W., Belgi, A., Espino, S.S., Olivera, B.M., Robinson, A.J., Chalmers, D.K., Norton, R.S.(2017) Toxicon 129: 113-122
- PubMed: 28216409 Search on PubMed
- DOI: https://doi.org/10.1016/j.toxicon.2017.02.012
- Primary Citation Related Structures:
5L34 - PubMed Abstract:
In natural proteins and peptides, amino acids exist almost invariably as l-isomers. There are, however, several examples of naturally-occurring peptides containing d-amino acids. In this study we investigated the role of a naturally-occurring d-amino acid in a small peptide identified in the transcriptome of a marine cone snail. This peptide belongs to a family of peptides known as contryphans, all of which contain a single d-amino acid residue. The solution structure of this peptide was solved by NMR, but further investigations with molecular dynamics simulations suggest that its solution behaviour may be more dynamic than suggested by the NMR ensemble. Functional tests in mice uncovered a novel bioactivity, a depressive phenotype that contrasts with the hyperactive phenotypes typically induced by contryphans. Trp3 is important for bioactivity, but this role is independent of the chirality at this position. The d-chirality of Trp3 in this peptide was found to be protective against enzymatic degradation. Analysis by NMR and molecular dynamics simulations indicated an interaction of Trp3 with lipid membranes, suggesting the possibility of a membrane-mediated mechanism of action for this peptide.
- Medicinal Chemistry, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville 3052, Victoria, Australia.
Organizational Affiliation: