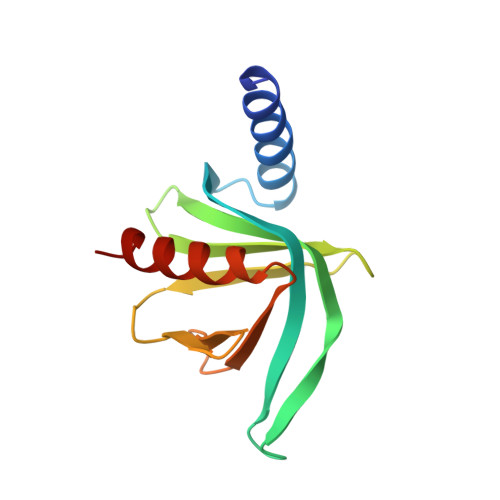
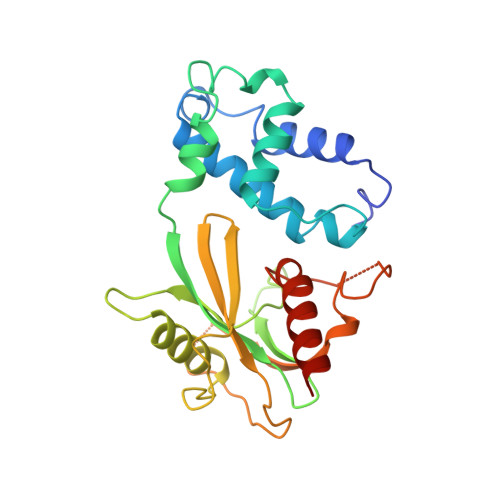
Structural basis of mRNA-cap recognition by Dcp1-Dcp2.
Mugridge, J.S., Ziemniak, M., Jemielity, J., Gross, J.D.(2016) Nat Struct Mol Biol 23: 987-994
- PubMed: 27694842 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.3301
- Primary Citation Related Structures:
5KQ1, 5KQ4 - PubMed Abstract:
Removal of the 5' cap on mRNA by the decapping enzyme Dcp2 is a critical step in 5'-to-3' mRNA decay. Understanding the structural basis of Dcp2 activity has been a challenge because Dcp2 is dynamic and has weak affinity for the cap substrate. Here we present a 2.6-Å-resolution crystal structure of a heterotrimer of fission yeast Dcp2, its essential activator Dcp1, and the human NMD cofactor PNRC2, in complex with a tight-binding cap analog. Cap binding is accompanied by a conformational change in Dcp2, thereby forming a composite nucleotide-binding site comprising conserved residues in the catalytic and regulatory domains. Kinetic analysis of PNRC2 revealed that a conserved short linear motif enhances both substrate affinity and the catalytic step of decapping. These findings explain why Dcp2 requires a conformational change for efficient catalysis and reveals that coactivators promote RNA binding and the catalytic step of decapping, possibly through different conformational states.
- Department of Pharmaceutical Chemistry, University of California, San Francisco, San Francisco, California, USA.
Organizational Affiliation: