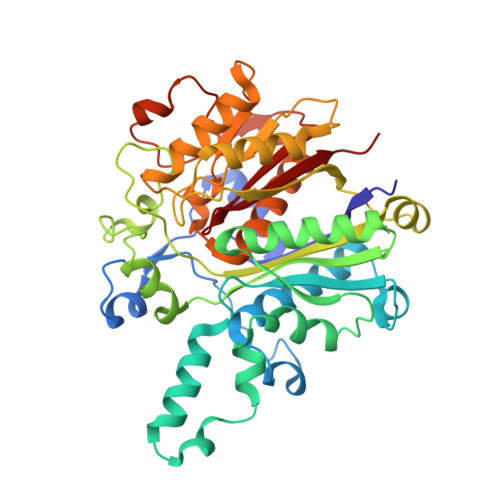
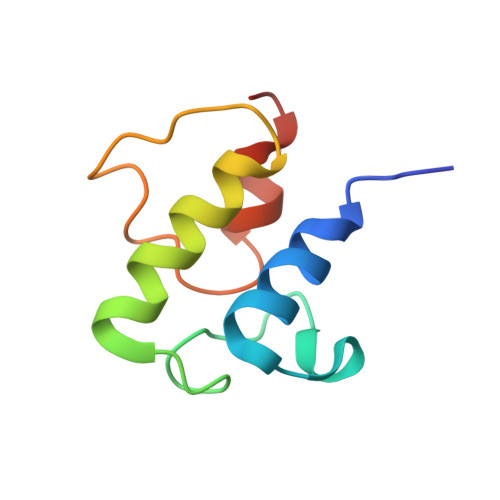
Molecular basis for interactions between an acyl carrier protein and a ketosynthase.
Milligan, J.C., Lee, D.J., Jackson, D.R., Schaub, A.J., Beld, J., Barajas, J.F., Hale, J.J., Luo, R., Burkart, M.D., Tsai, S.C.(2019) Nat Chem Biol 15: 669-671
- PubMed: 31209348 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-019-0301-y
- Primary Citation Related Structures:
5KOF - PubMed Abstract:
Fatty acid synthases are dynamic ensembles of enzymes that can biosynthesize long hydrocarbon chains efficiently. Here we visualize the interaction between the Escherichia coli acyl carrier protein (AcpP) and β-ketoacyl-ACP-synthase I (FabB) using X-ray crystallography, NMR, and molecular dynamics simulations. We leveraged this structural information to alter lipid profiles in vivo and provide a molecular basis for how protein-protein interactions can regulate the fatty acid profile in E. coli.
- Department of Molecular Biology and Biochemistry, University of California, Irvine, Irvine, CA, USA.
Organizational Affiliation: