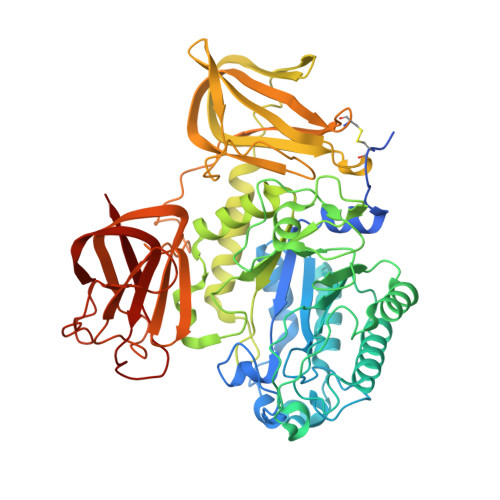
The structure of a glycoside hydrolase 29 family member from a rumen bacterium reveals unique, dual carbohydrate-binding domains.
Summers, E.L., Moon, C.D., Atua, R., Arcus, V.L.(2016) Acta Crystallogr F Struct Biol Commun 72: 750-761
- PubMed: 27710940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16014072
- Primary Citation Related Structures:
5K9H - PubMed Abstract:
Glycoside hydrolase (GH) family 29 consists solely of α-L-fucosidases. These enzymes catalyse the hydrolysis of glycosidic bonds. Here, the structure of GH29_0940, a protein cloned from metagenomic DNA from the rumen of a cow, has been solved, which reveals a multi-domain arrangement that has only recently been identified in bacterial GH29 enzymes. The microbial species that provided the source of this enzyme is unknown. This enzyme contains a second carbohydrate-binding domain at its C-terminal end in addition to the typical N-terminal catalytic domain and carbohydrate-binding domain arrangement of GH29-family proteins. GH29_0940 is a monomer and its overall structure consists of an N-terminal TIM-barrel-like domain, a central β-sandwich domain and a C-terminal β-sandwich domain. The TIM-barrel-like catalytic domain exhibits a (β/α) 8/7 arrangement in the core instead of the typical (β/α) 8 topology, with the `missing' α-helix replaced by a long meandering loop that `closes' the barrel structure and suggests a high degree of structural flexibility in the catalytic core. This feature was also noted in all six other structures of GH29 enzymes that have been deposited in the PDB. Based on sequence and structural similarity, the residues Asp162 and Glu220 are proposed to serve as the catalytic nucleophile and the proton donor, respectively. Like other GH29 enzymes, the GH29_0940 structure shows five strictly conserved residues in the catalytic pocket. The structure shows two glycerol molecules in the active site, which have also been observed in other GH29 structures, suggesting that the enzyme catalyses the hydrolysis of small carbohydrates. The two binding domains are classed as family 32 carbohydrate-binding modules (CBM32). These domains have residues involved in ligand binding in the loop regions at the edge of the β-sandwich. The predicted substrate-binding residues differ between the modules, suggesting that different modules bind to different groups on the substrate(s). Enzymes that possess multiple copies of CBMs are thought to have a complex mechanism of ligand recognition. Defined electron density identifying a long 20-amino-acid hydrophilic loop separating the two CBMs was observed. This suggests that the additional C-terminal domain may have a dynamic range of movement enabled by the loop, allowing a unique mode of action for a GH29 enzyme that has not been identified previously.
- Biological Sciences, The University of Waikato, Private Bag 3105, Hamilton 3240, New Zealand.
Organizational Affiliation: