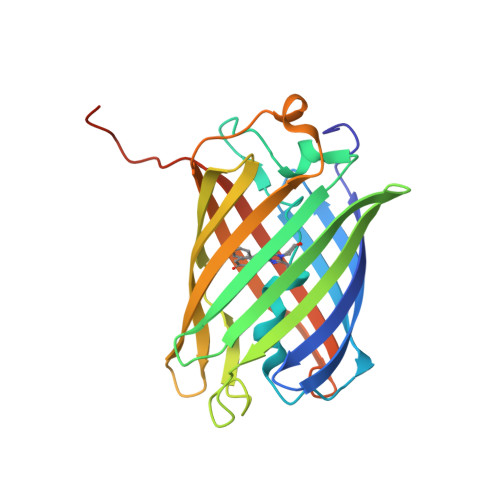
The crystal structure of red fluorescent protein TagRFP-T reveals the mechanism of its superior photostability.
Liu, R., Liang, Q.N., Du, S.Q., Hu, X.J., Ding, Y.(2016) Biochem Biophys Res Commun 477: 229-234
- PubMed: 27297107 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2016.06.047
- Primary Citation Related Structures:
5JVA - PubMed Abstract:
The red fluorescent protein variant TagRFP-T has greatly improved photostability over its parent molecule, TagRFP, but the underlying mechanism leading to this improvement is to date unknown. The 1.95 Å resolution crystallographic structure of TagRFP-T showed that its chromophore exists as a mixture of cis and trans coplanar isomers in roughly equal proportions. Interestingly, both isomers are able to fluoresce, a property that has never been observed in any other fluorescent protein. We propose a "circular restoration model" for TagRFP-T to explain its superior photostability: There are four co-existing chromophore states (cis/trans protonated/ionized state) that can be driven by light to transform from one state into another. This model also explains how TagRPF-T essentially eliminates the temporary dark state (reversible photobleaching).
- Department of Physiology and Biophysics, School of Life Sciences, Fudan University, Shanghai 200438, China.
Organizational Affiliation: