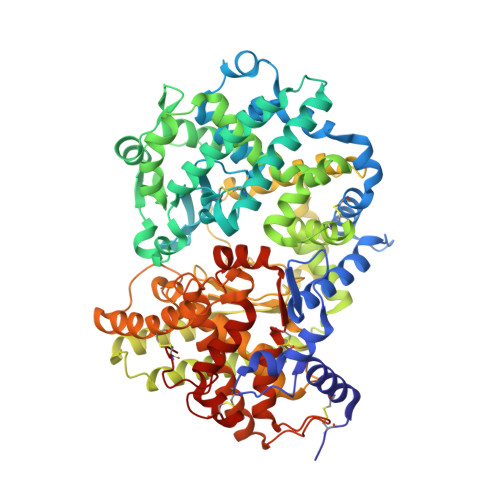
Structure of neprilysin in complex with the active metabolite of sacubitril.
Schiering, N., D'Arcy, A., Villard, F., Ramage, P., Logel, C., Cumin, F., Ksander, G.M., Wiesmann, C., Karki, R.G., Mogi, M.(2016) Sci Rep 6: 27909-27909
- PubMed: 27302413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep27909
- Primary Citation Related Structures:
5JMY - PubMed Abstract:
Sacubitril is an ethyl ester prodrug of LBQ657, the active neprilysin (NEP) inhibitor, and a component of LCZ696 (sacubitril/valsartan). We report herein the three-dimensional structure of LBQ657 in complex with human NEP at 2 Å resolution. The crystal structure unravels the binding mode of the compound occupying the S1, S1' and S2' sub-pockets of the active site, consistent with a competitive inhibition mode. An induced fit conformational change upon binding of the P1'-biphenyl moiety of the inhibitor suggests an explanation for its selectivity against structurally homologous zinc metallopeptidases.
- Novartis Institutes for BioMedical Research Inc., Fabrikstrasse 16, CH-4002 Basel, Switzerland.
Organizational Affiliation: