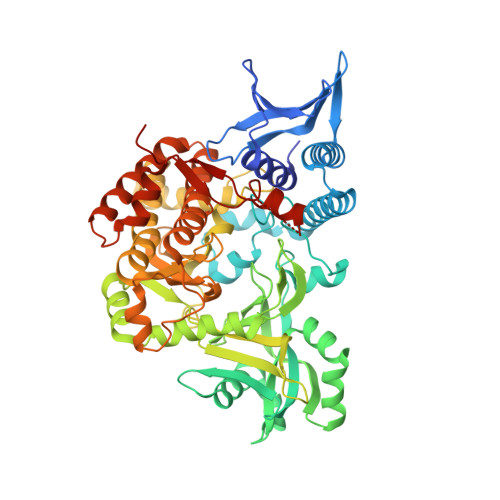
Structural and Functional Characterization of Aerobactin Synthetase IucA from a Hypervirulent Pathotype of Klebsiella pneumoniae.
Bailey, D.C., Drake, E.J., Grant, T.D., Gulick, A.M.(2016) Biochemistry 55: 3559-3570
- PubMed: 27253399 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b00409
- Primary Citation Related Structures:
5JM7, 5JM8 - PubMed Abstract:
Iron is a vital mineral nutrient required by virtually all life forms to prosper; pathogenic bacteria are no exception. Despite the abundance of iron within the human host, highly regulated iron physiology can result in exceedingly low levels of iron bioavailable to prospective invading bacteria. To combat this scarcity of iron, many pathogenic bacteria have acquired specific and efficient iron acquisition systems, which allow them to thrive in iron-deficient host environments. One of the more prominent bacterial iron acquisition systems involves the synthesis, secretion, and reuptake of small-molecule iron chelators known as siderophores. Aerobactin, a citrate-hydroxamate siderophore originally isolated nearly 50 years ago, is produced by a number of pathogenic Gram-negative bacteria. Aerobactin has recently been demonstrated to play a pivotal role in mediating the enhanced virulence of a particularly invasive pathotype of Klebsiella pneumoniae (hvKP). Toward further understanding of this key virulence factor, we report the structural and functional characterization of aerobactin synthetase IucA from a strain of hvKP. The X-ray crystal structures of unliganded and ATP-bound forms of IucA were solved, forming the foundation of our structural analysis. Small angle X-ray scattering (SAXS) data suggest that, unlike its closest structurally characterized homologues, IucA adopts a tetrameric assembly in solution. Finally, we employed activity assays to investigate the substrate specificity and determine the apparent steady-state kinetic parameters of IucA.
- The Hauptman-Woodward Medical Research Institute , Buffalo, New York, United States.
Organizational Affiliation: