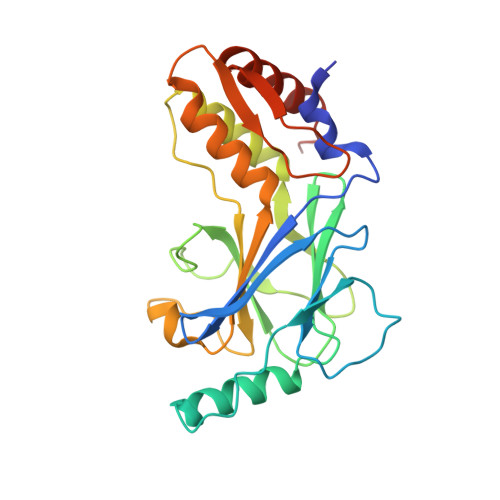
Structural basis for concerted recruitment and activation of IRF-3 by innate immune adaptor proteins.
Zhao, B., Shu, C., Gao, X., Sankaran, B., Du, F., Shelton, C.L., Herr, A.B., Ji, J.Y., Li, P.(2016) Proc Natl Acad Sci U S A 113: E3403-E3412
- PubMed: 27302953 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1603269113
- Primary Citation Related Structures:
5JEJ, 5JEK, 5JEL, 5JEM, 5JEO, 5JER - PubMed Abstract:
Type I IFNs are key cytokines mediating innate antiviral immunity. cGMP-AMP synthase, ritinoic acid-inducible protein 1 (RIG-I)-like receptors, and Toll-like receptors recognize microbial double-stranded (ds)DNA, dsRNA, and LPS to induce the expression of type I IFNs. These signaling pathways converge at the recruitment and activation of the transcription factor IRF-3 (IFN regulatory factor 3). The adaptor proteins STING (stimulator of IFN genes), MAVS (mitochondrial antiviral signaling), and TRIF (TIR domain-containing adaptor inducing IFN-β) mediate the recruitment of IRF-3 through a conserved pLxIS motif. Here we show that the pLxIS motif of phosphorylated STING, MAVS, and TRIF binds to IRF-3 in a similar manner, whereas residues upstream of the motif confer specificity. The structure of the IRF-3 phosphomimetic mutant S386/396E bound to the cAMP response element binding protein (CREB)-binding protein reveals that the pLxIS motif also mediates IRF-3 dimerization and activation. Moreover, rotavirus NSP1 (nonstructural protein 1) employs a pLxIS motif to target IRF-3 for degradation, but phosphorylation of NSP1 is not required for its activity. These results suggest a concerted mechanism for the recruitment and activation of IRF-3 that can be subverted by viral proteins to evade innate immune responses.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX 77843;
Organizational Affiliation: