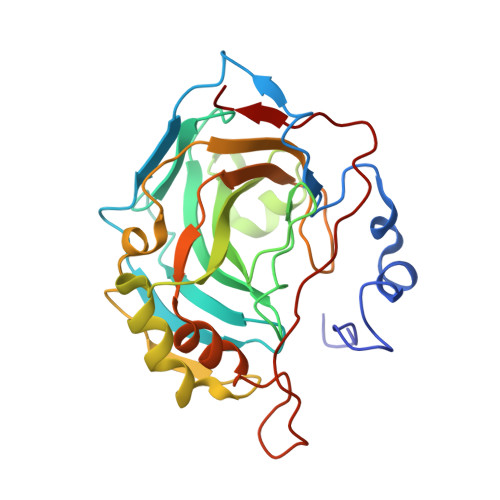
Kinetic and X-ray crystallographic investigations of substituted 2-thio-6-oxo-1,6-dihydropyrimidine-benzenesulfonamides acting as carbonic anhydrase inhibitors.
Vullo, D., Supuran, C.T., Scozzafava, A., De Simone, G., Monti, S.M., Alterio, V., Carta, F.(2016) Bioorg Med Chem 24: 3643-3648
- PubMed: 27316543 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2016.06.005
- Primary Citation Related Structures:
5J8Z - PubMed Abstract:
Herein we report an in vitro kinetic evaluation against the most relevant human carbonic anhydrase (hCA, EC 4.2.1.1) isoforms (I, II, IX and XII) of a small series of lactate dehydrogenase (LDH, EC 1.1.1.27) inhibitors. All compounds contain a primary sulfonamide zinc-binding group (ZBG) substituted with the 2-thio-6-oxo-1,6-dihydropyrimidine scaffold. By means of X-ray crystallographic experiments we explored the ligand-enzyme binding modes, thus highlighting the contribution of the 2-thio-6-oxo-1,6-dihydropyrimidine moiety to the stabilization of the complex.
- Università degli Studi di Firenze, Dipartimento di Chimica, Laboratorio di Chimica Bioinorganica, Rm. 188, Via della Lastruccia 3, 50019 Sesto Fiorentino (Florence), Italy.
Organizational Affiliation: