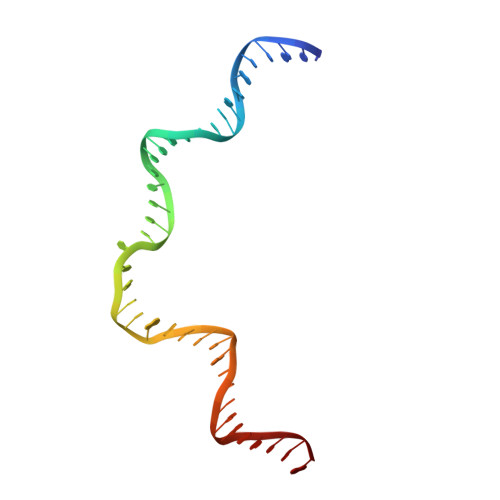
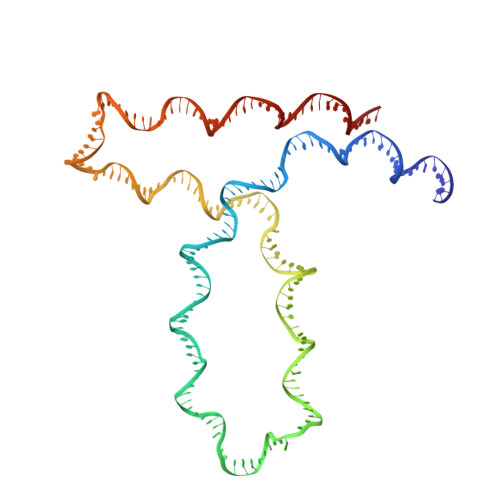
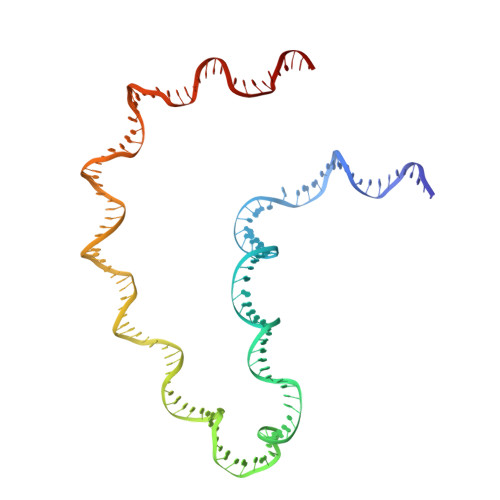
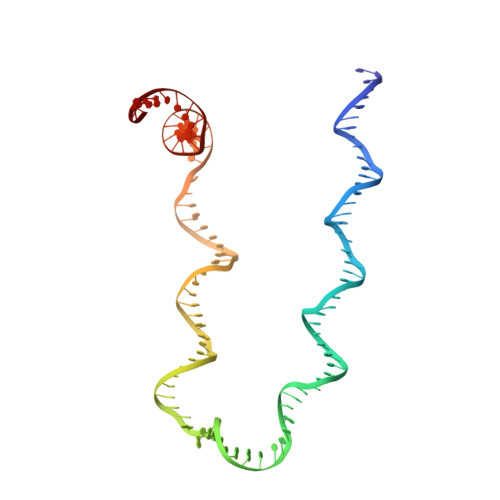
Structure of a Holliday junction complex reveals mechanisms governing a highly regulated DNA transaction.
Laxmikanthan, G., Xu, C., Brilot, A.F., Warren, D., Steele, L., Seah, N., Tong, W., Grigorieff, N., Landy, A., Van Duyne, G.D.(2016) Elife 5
- PubMed: 27223329 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.14313
- Primary Citation Related Structures:
5J0N - PubMed Abstract:
The molecular machinery responsible for DNA expression, recombination, and compaction has been difficult to visualize as functionally complete entities due to their combinatorial and structural complexity. We report here the structure of the intact functional assembly responsible for regulating and executing a site-specific DNA recombination reaction. The assembly is a 240-bp Holliday junction (HJ) bound specifically by 11 protein subunits. This higher-order complex is a key intermediate in the tightly regulated pathway for the excision of bacteriophage λ viral DNA out of the E. coli host chromosome, an extensively studied paradigmatic model system for the regulated rearrangement of DNA. Our results provide a structural basis for pre-existing data describing the excisive and integrative recombination pathways, and they help explain their regulation.
- Department of Molecular Biology, Cell Biology, and Biochemistry, Brown University, Providence, United States.
Organizational Affiliation: