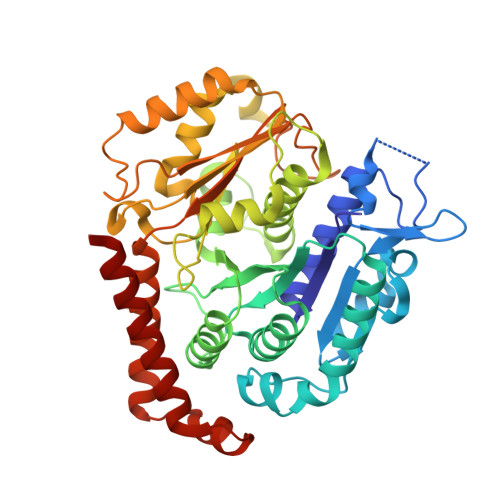
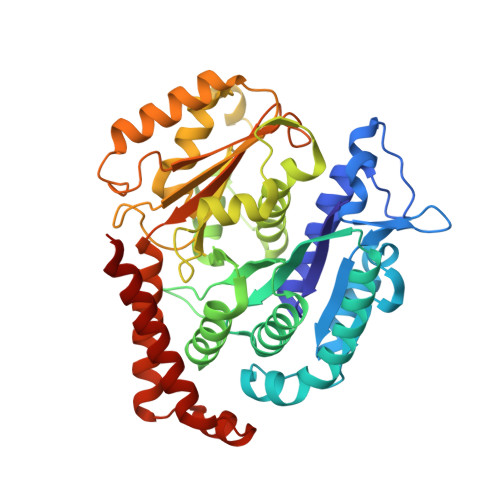
Mutations in Human Tubulin Proximal to the Kinesin-Binding Site Alter Dynamic Instability at Microtubule Plus- and Minus-Ends.
Ti, S.C., Pamula, M.C., Howes, S.C., Duellberg, C., Cade, N.I., Kleiner, R.E., Forth, S., Surrey, T., Nogales, E., Kapoor, T.M.(2016) Dev Cell 37: 72-84
- PubMed: 27046833 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.devcel.2016.03.003
- Primary Citation Related Structures:
5IJ0, 5IJ9 - PubMed Abstract:
The assembly of microtubule-based cellular structures depends on regulated tubulin polymerization and directional transport. Here, we purify and characterize tubulin heterodimers that have human β-tubulin isotype III (TUBB3), as well as heterodimers with one of two β-tubulin mutations (D417H or R262H). Both point mutations are proximal to the kinesin-binding site and have been linked to an ocular motility disorder in humans. Compared to wild-type, microtubules with these mutations have decreased catastrophe frequencies and increased average lifetimes of plus- and minus-end-stabilizing caps. Importantly, the D417H mutation does not alter microtubule lattice structure or Mal3 binding to growing filaments. Instead, this mutation reduces the affinity of tubulin for TOG domains and colchicine, suggesting that the distribution of tubulin heterodimer conformations is changed. Together, our findings reveal how residues on the surface of microtubules, distal from the GTP-hydrolysis site and inter-subunit contacts, can alter polymerization dynamics at the plus- and minus-ends of microtubules.
- Laboratory of Chemistry and Cell Biology, The Rockefeller University, 1230 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: