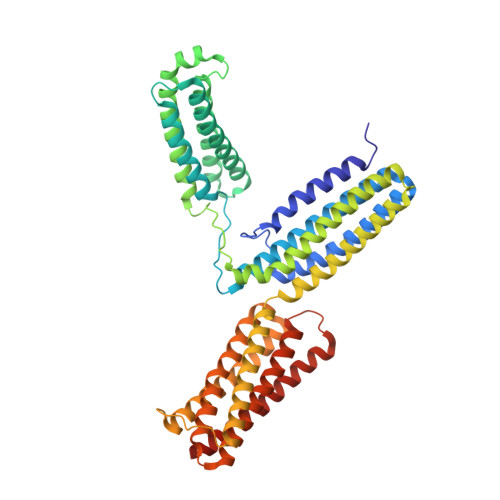
Structural and Functional Analysis of a Talin Triple-Domain Module Suggests an Alternative Talin Autoinhibitory Configuration.
Zhang, H., Chang, Y.C., Huang, Q., Brennan, M.L., Wu, J.(2016) Structure 24: 721-729
- PubMed: 27150043 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.02.020
- Primary Citation Related Structures:
5IC0, 5IC1 - PubMed Abstract:
Talin plays an important role in regulating integrin-mediated signaling. Talin function is autoinhibited by intramolecular interactions between the integrin-binding F3 domain and the autoinhibitory domain (R9). We determined the crystal structure of a triple-domain fragment, R7R8R9, which contains R9 and the RIAM (Rap1-interacting adaptor molecule) binding domain (R8). The structure reveals a crystallographic contact between R9 and a symmetrically related R8 domain, representing a homodimeric interaction in talin. Strikingly, we demonstrated that the α5 helix of R9 also interacts with the F3 domain, despite no interdomain contact involving the α5 helix in the crystal structure of an F2F3:R9 autoinhibitory complex reported previously. Mutations on the α5 helix significantly diminish the F3:R9 association and lead to elevated talin activity. Our results offer biochemical and functional evidence of the existence of a new talin autoinhibitory configuration, thus providing a more comprehensive understanding of talin autoinhibition, regulation, and quaternary structure assembly.
- Molecular Therapeutics Program, Fox Chase Cancer Center, Philadelphia, PA 19111, USA.
Organizational Affiliation: