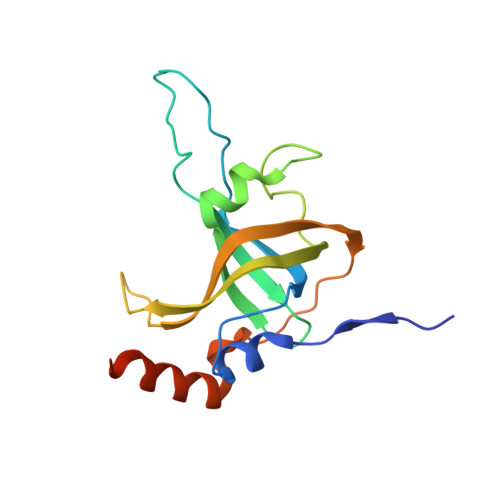
Structural and functional consequences of a disease mutation in the telomere protein TPP1.
Bisht, K., Smith, E.M., Tesmer, V.M., Nandakumar, J.(2016) Proc Natl Acad Sci U S A 113: 13021-13026
- PubMed: 27807141 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1605685113
- Primary Citation Related Structures:
5I2X, 5I2Y - PubMed Abstract:
Telomerase replicates chromosome ends to facilitate continued cell division. Mutations that compromise telomerase function result in stem cell failure diseases, such as dyskeratosis congenita (DC). One such mutation (K170Δ), residing in the telomerase-recruitment factor TPP1, provides an excellent opportunity to structurally, biochemically, and genetically dissect the mechanism of such diseases. We show through site-directed mutagenesis and X-ray crystallography that this TPP1 disease mutation deforms the conformation of two critical amino acids of the TEL [TPP1's glutamate (E) and leucine-rich (L)] patch, the surface of TPP1 that binds telomerase. Using CRISPR-Cas9 technology, we demonstrate that introduction of this mutation in a heterozygous manner is sufficient to shorten telomeres in human cells. Our findings rule out dominant-negative effects of the mutation. Instead, these findings implicate reduced TEL patch dosage in causing telomere shortening. Our studies provide mechanistic insight into telomerase-deficiency diseases and encourage the development of gene therapies to counter such diseases.
- Department of Molecular, Cellular, and Developmental Biology, University of Michigan, Ann Arbor, MI 48109.
Organizational Affiliation: