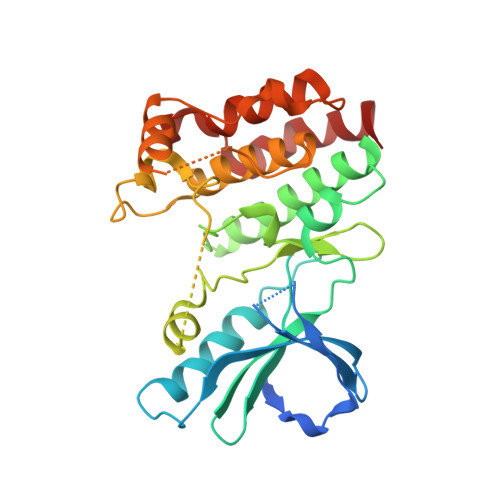
DNA-Encoded Library Screening Identifies Benzo[b][1,4]oxazepin-4-ones as Highly Potent and Monoselective Receptor Interacting Protein 1 Kinase Inhibitors.
Harris, P.A., King, B.W., Bandyopadhyay, D., Berger, S.B., Campobasso, N., Capriotti, C.A., Cox, J.A., Dare, L., Dong, X., Finger, J.N., Grady, L.C., Hoffman, S.J., Jeong, J.U., Kang, J., Kasparcova, V., Lakdawala, A.S., Lehr, R., McNulty, D.E., Nagilla, R., Ouellette, M.T., Pao, C.S., Rendina, A.R., Schaeffer, M.C., Summerfield, J.D., Swift, B.A., Totoritis, R.D., Ward, P., Zhang, A., Zhang, D., Marquis, R.W., Bertin, J., Gough, P.J.(2016) J Med Chem 59: 2163-2178
- PubMed: 26854747 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01898
- Primary Citation Related Structures:
5HX6 - PubMed Abstract:
The recent discovery of the role of receptor interacting protein 1 (RIP1) kinase in tumor necrosis factor (TNF)-mediated inflammation has led to its emergence as a highly promising target for the treatment of multiple inflammatory diseases. We screened RIP1 against GSK's DNA-encoded small-molecule libraries and identified a novel highly potent benzoxazepinone inhibitor series. We demonstrate that this template possesses complete monokinase selectivity for RIP1 plus unique species selectivity for primate versus nonprimate RIP1. We elucidate the conformation of RIP1 bound to this benzoxazepinone inhibitor driving its high kinase selectivity and design specific mutations in murine RIP1 to restore potency to levels similar to primate RIP1. This series differentiates itself from known RIP1 inhibitors in combining high potency and kinase selectivity with good pharmacokinetic profiles in rodents. The favorable developability profile of this benzoxazepinone template, as exemplified by compound 14 (GSK'481), makes it an excellent starting point for further optimization into a RIP1 clinical candidate.
- Platform Technology & Science, GlaxoSmithKline , Winter Street, Waltham, Massachusetts 02451, United States.
Organizational Affiliation: