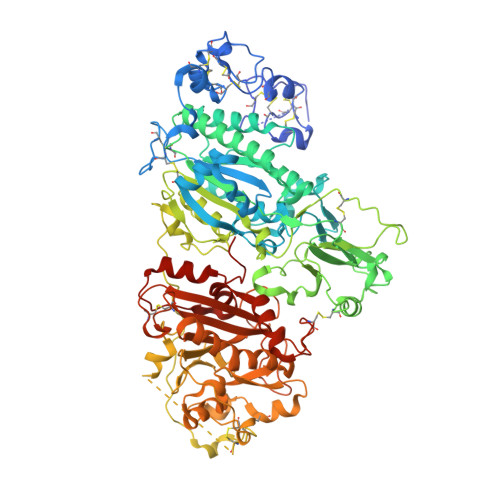
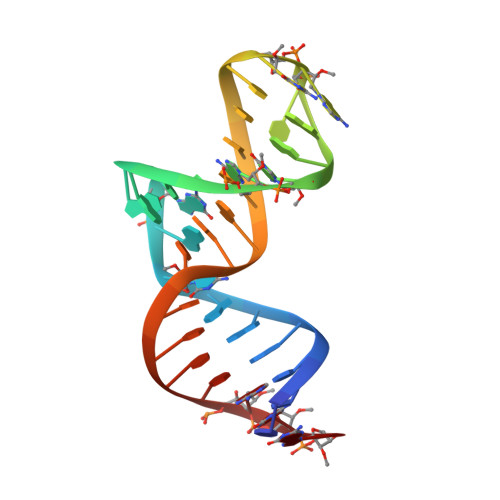
Structural basis for specific inhibition of Autotaxin by a DNA aptamer
Kato, K., Ikeda, H., Miyakawa, S., Futakawa, S., Nonaka, Y., Fujiwara, M., Okudaira, S., Kano, K., Aoki, J., Morita, J., Ishitani, R., Nishimasu, H., Nakamura, Y., Nureki, O.(2016) Nat Struct Mol Biol 23: 395-401
- PubMed: 27043297 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3200
- Primary Citation Related Structures:
5HRT - PubMed Abstract:
ATX is a plasma lysophospholipase D that hydrolyzes lysophosphatidylcholine (LPC) and produces lysophosphatidic acid. To date, no ATX-inhibition-mediated treatment strategies for human diseases have been established. Here, we report anti-ATX DNA aptamers that inhibit ATX with high specificity and efficacy. We solved the crystal structure of ATX in complex with the anti-ATX aptamer RB011, at 2.0-Å resolution. RB011 binds in the vicinity of the active site through base-specific interactions, thus preventing the access of the choline moiety of LPC substrates. Using the structural information, we developed the modified anti-ATX DNA aptamer RB014, which exhibited in vivo efficacy in a bleomycin-induced pulmonary fibrosis mouse model. Our findings reveal the structural basis for the specific inhibition of ATX by the anti-ATX aptamer and highlight the therapeutic potential of anti-ATX aptamers for the treatment of human diseases, such as pulmonary fibrosis.
- Department of Biological Sciences, Graduate School of Science, University of Tokyo, Tokyo, Japan.
Organizational Affiliation: