System-Wide Modulation of HECT E3 Ligases with Selective Ubiquitin Variant Probes.
Zhang, W., Wu, K.P., Sartori, M.A., Kamadurai, H.B., Ordureau, A., Jiang, C., Mercredi, P.Y., Murchie, R., Hu, J., Persaud, A., Mukherjee, M., Li, N., Doye, A., Walker, J.R., Sheng, Y., Hao, Z., Li, Y., Brown, K.R., Lemichez, E., Chen, J., Tong, Y., Harper, J.W., Moffat, J., Rotin, D., Schulman, B.A., Sidhu, S.S.(2016) Mol Cell 62: 121-136
- PubMed: 26949039 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2016.02.005
- Primary Citation Related Structures:
5C7J, 5C7M, 5HPK, 5HPL, 5HPS, 5HPT - PubMed Abstract:
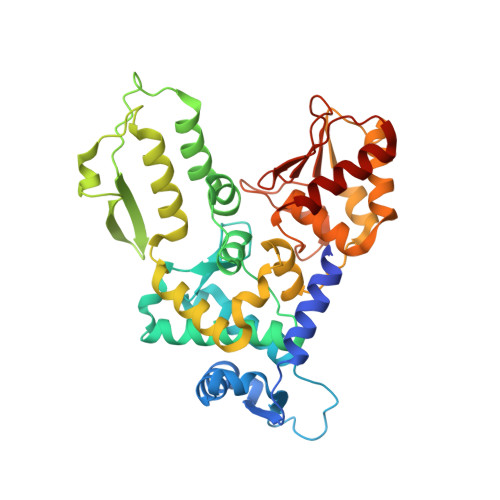
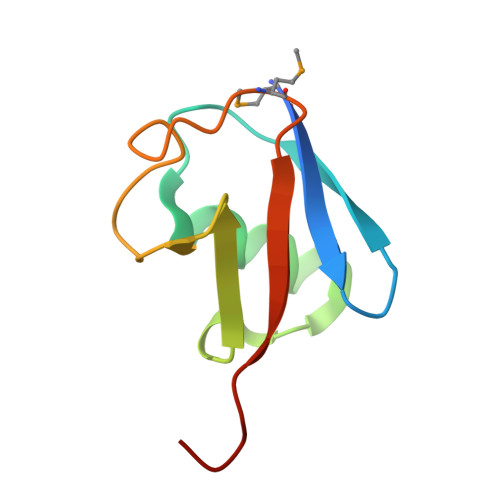
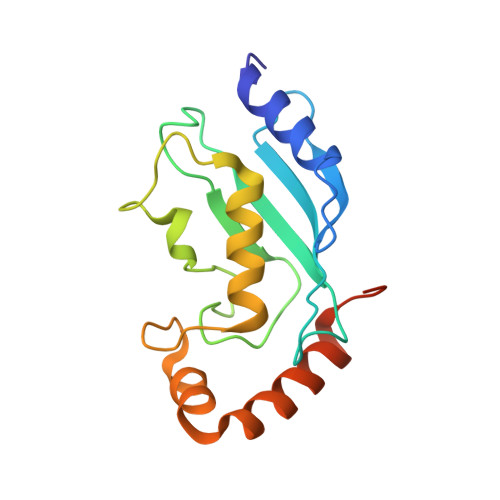
HECT-family E3 ligases ubiquitinate protein substrates to control virtually every eukaryotic process and are misregulated in numerous diseases. Nonetheless, understanding of HECT E3s is limited by a paucity of selective and potent modulators. To overcome this challenge, we systematically developed ubiquitin variants (UbVs) that inhibit or activate HECT E3s. Structural analysis of 6 HECT-UbV complexes revealed UbV inhibitors hijacking the E2-binding site and activators occupying a ubiquitin-binding exosite. Furthermore, UbVs unearthed distinct regulation mechanisms among NEDD4 subfamily HECTs and proved useful for modulating therapeutically relevant targets of HECT E3s in cells and intestinal organoids, and in a genetic screen that identified a role for NEDD4L in regulating cell migration. Our work demonstrates versatility of UbVs for modulating activity across an E3 family, defines mechanisms and provides a toolkit for probing functions of HECT E3s, and establishes a general strategy for systematic development of modulators targeting families of signaling proteins.
- Donnelly Centre for Cellular and Biomolecular Research, Banting and Best Department of Medical Research, University of Toronto, 160 College Street, Toronto, ON M5S3E1, Canada.
Organizational Affiliation: