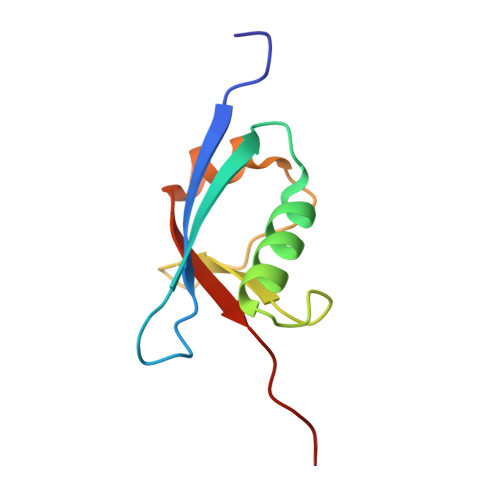
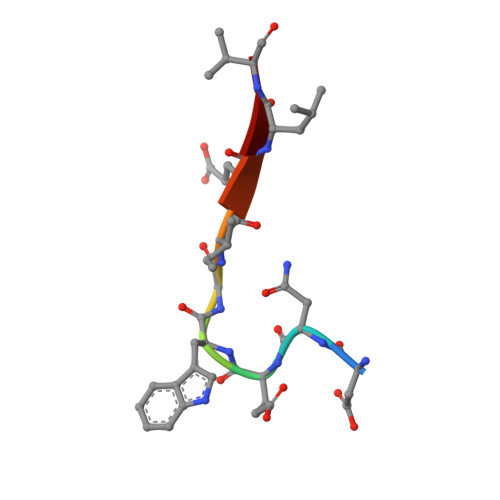
Structural and Functional Analysis of a Novel Interaction Motif within UFM1-activating Enzyme 5 (UBA5) Required for Binding to Ubiquitin-like Proteins and Ufmylation.
Habisov, S., Huber, J., Ichimura, Y., Akutsu, M., Rogova, N., Loehr, F., McEwan, D.G., Johansen, T., Dikic, I., Doetsch, V., Komatsu, M., Rogov, V.V., Kirkin, V.(2016) J Biological Chem 291: 9025-9041
- PubMed: 26929408 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.715474
- Primary Citation Related Structures:
5HKH - PubMed Abstract:
The covalent conjugation of ubiquitin-fold modifier 1 (UFM1) to proteins generates a signal that regulates transcription, response to cell stress, and differentiation. Ufmylation is initiated by ubiquitin-like modifier activating enzyme 5 (UBA5), which activates and transfers UFM1 to ubiquitin-fold modifier-conjugating enzyme 1 (UFC1). The details of the interaction between UFM1 and UBA5 required for UFM1 activation and its downstream transfer are however unclear. In this study, we described and characterized a combined linear LC3-interacting region/UFM1-interacting motif (LIR/UFIM) within the C terminus of UBA5. This single motif ensures that UBA5 binds both UFM1 and light chain 3/γ-aminobutyric acid receptor-associated proteins (LC3/GABARAP), two ubiquitin (Ub)-like proteins. We demonstrated that LIR/UFIM is required for the full biological activity of UBA5 and for the effective transfer of UFM1 onto UFC1 and a downstream protein substrate both in vitro and in cells. Taken together, our study provides important structural and functional insights into the interaction between UBA5 and Ub-like modifiers, improving the understanding of the biology of the ufmylation pathway.
- From Translational Innovation Platform Oncology, Merck KGaA, Frankfurter Strasse 250, 64293 Darmstadt, Germany, the Institute of Biochemistry II, Goethe University School of Medicine, Theodor-Stern-Kai 7, 60590 Frankfurt am Main, Germany, the BMLS Buchmann Institute for Molecular Life Sciences, Goethe University Frankfurt, Riedberg Campus, Max-von-Laue-Strasse 15, 60438 Frankfurt am Main, Germany.
Organizational Affiliation: