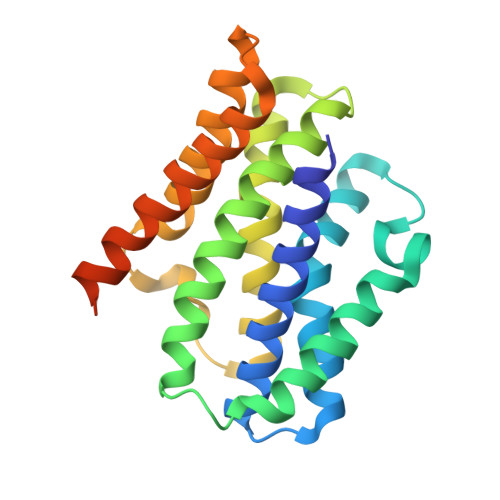
Crystal structures of the TRIC trimeric intracellular cation channel orthologues
Kasuya, G., Hiraizumi, M., Maturana, A.D., Kumazaki, K., Fujiwara, Y., Liu, K., Nakada-Nakura, Y., Iwata, S., Tsukada, K., Komori, T., Uemura, S., Goto, Y., Nakane, T., Takemoto, M., Kato, H.E., Yamashita, K., Wada, M., Ito, K., Ishitani, R., Hattori, M., Nureki, O.(2016) Cell Res 26: 1288-1301
- PubMed: 27909292 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cr.2016.140
- Primary Citation Related Structures:
5H35, 5H36 - PubMed Abstract:
Ca 2+ release from the sarcoplasmic reticulum (SR) and endoplasmic reticulum (ER) is crucial for muscle contraction, cell growth, apoptosis, learning and memory. The trimeric intracellular cation (TRIC) channels were recently identified as cation channels balancing the SR and ER membrane potentials, and are implicated in Ca 2+ signaling and homeostasis. Here we present the crystal structures of prokaryotic TRIC channels in the closed state and structure-based functional analyses of prokaryotic and eukaryotic TRIC channels. Each trimer subunit consists of seven transmembrane (TM) helices with two inverted repeated regions. The electrophysiological, biochemical and biophysical analyses revealed that TRIC channels possess an ion-conducting pore within each subunit, and that the trimer formation contributes to the stability of the protein. The symmetrically related TM2 and TM5 helices are kinked at the conserved glycine clusters, and these kinks are important for the channel activity. Furthermore, the kinks of the TM2 and TM5 helices generate lateral fenestrations at each subunit interface. Unexpectedly, these lateral fenestrations are occupied with lipid molecules. This study provides the structural and functional framework for the molecular mechanism of this ion channel superfamily.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, Tokyo 113-0032, Japan.
Organizational Affiliation: