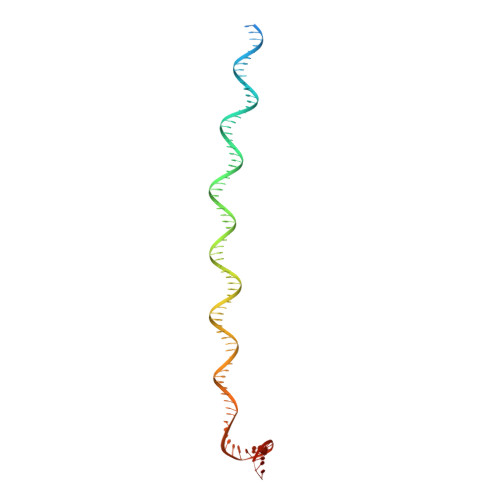
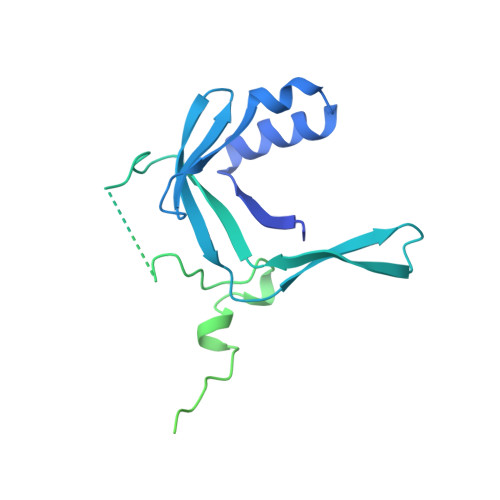
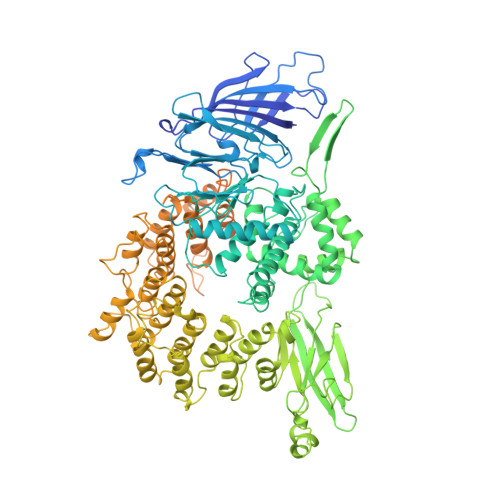
Structure of Promoter-Bound TFIID and Model of Human Pre-Initiation Complex Assembly.
Louder, R.K., He, Y., Lopez-Blanco, J.R., Fang, J., Chacon, P., Nogales, E.(2016) Nature 531: 604
- PubMed: 27007846 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature17394
- Primary Citation Related Structures:
5FUR - PubMed Abstract:
The general transcription factor IID (TFIID) plays a central role in the initiation of RNA polymerase II (Pol II)-dependent transcription by nucleating pre-initiation complex (PIC) assembly at the core promoter. TFIID comprises the TATA-binding protein (TBP) and 13 TBP-associated factors (TAF1-13), which specifically interact with a variety of core promoter DNA sequences. Here we present the structure of human TFIID in complex with TFIIA and core promoter DNA, determined by single-particle cryo-electron microscopy at sub-nanometre resolution. All core promoter elements are contacted by subunits of TFIID, with TAF1 and TAF2 mediating major interactions with the downstream promoter. TFIIA bridges the TBP-TATA complex with lobe B of TFIID. We also present the cryo-electron microscopy reconstruction of a fully assembled human TAF-less PIC. Superposition of common elements between the two structures provides novel insights into the general role of TFIID in promoter recognition, PIC assembly, and transcription initiation.
- Biophysics Graduate Group, University of California, Berkeley, California 94720, USA.
Organizational Affiliation: