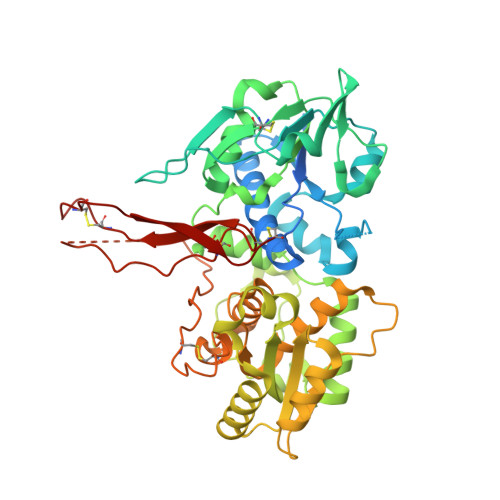
A Proactive Role of Water Molecules in Acceptor Recognition of Thrombospondin Type 1 Repeats by Protein-O-Fucosyltransferase 2
Valero-Gonzalez, J., Leonhard-Melief, C., Lira-Navarrete, E., Jimenez-Oses, G., Hernandez-Ruiz, C., Pallares, M.C., Yruela, I., Vasudevan, D., Lostao, A., Corzana, F., Takeuchi, H., Haltiwanger, R.S., Hurtado-Guerrero, R.(2016) Nat Chem Biol 12: 240
- PubMed: 26854667
- DOI: https://doi.org/10.1038/nchembio.2019
- Primary Citation of Related Structures:
5FOE - PubMed Abstract:
Protein O-fucosyltransferase 2 (POFUT2) is an essential enzyme that fucosylates serine and threonine residues of folded thrombospondin type 1 repeats (TSRs). To date, the mechanism by which this enzyme recognizes very dissimilar TSRs has been unclear. By engineering a fusion protein, we report the crystal structure of Caenorhabditis elegans POFUT2 (CePOFUT2) in complex with GDP and human TSR1 that suggests an inverting mechanism for fucose transfer assisted by a catalytic base and shows that nearly half of the TSR1 is embraced by CePOFUT2. A small number of direct interactions and a large network of water molecules maintain the complex. Site-directed mutagenesis demonstrates that POFUT2 fucosylates threonine preferentially over serine and relies on folded TSRs containing the minimal consensus sequence C-X-X-S/T-C. Crystallographic and mutagenesis data, together with atomic-level simulations, uncover a binding mechanism by which POFUT2 promiscuously recognizes the structural fingerprint of poorly homologous TSRs through a dynamic network of water-mediated interactions.
- Institute for Biocomputation and Physics of Complex Systems (BIFI), University of Zaragoza, BIFI-IQFR (CSIC) Joint Unit, Mariano Esquillor s/n, Campus Rio Ebro, Zaragoza, Spain.
Organizational Affiliation: