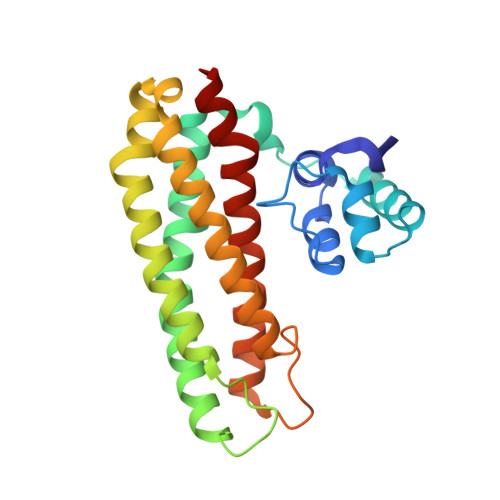
Crystal Structure of the Repair of Iron Centers Protein Ytfe and its Interaction with No
Lo, F.-C., Hsieh, C.-C., Maestre-Reyna, M., Chen, C.-Y., Ko, T.-P., Horng, Y.-C., Lai, Y.-C., Chiang, Y.-W., Chou, C.-M., Chiang, C.-H., Huang, W.-N., Liaw, W.-F.(2016) Chemistry 22: 9768
- PubMed: 27246459 Search on PubMed
- DOI: https://doi.org/10.1002/chem.201600990
- Primary Citation Related Structures:
5FNN, 5FNP, 5FNY - PubMed Abstract:
Molecular mechanisms underlying the repair of nitrosylated [Fe-S] clusters by the microbial protein YtfE remain poorly understood. The X-ray crystal structure of YtfE, in combination with EPR, magnetic circular dichroism (MCD), UV, and (17) O-labeling electron spin echo envelope modulation measurements, show that each iron of the oxo-bridged Fe(II) -Fe(III) diiron core is coordinatively unsaturated with each iron bound to two bridging carboxylates and two terminal histidines in addition to an oxo-bridge. Structural analysis reveals that there are two solvent-accessible tunnels, both of which converge to the diiron center and are critical for capturing substrates. The reactivity of the reduced-form Fe(II) -Fe(II) YtfE toward nitric oxide demonstrates that the prerequisite for N2 O production requires the two iron sites to be nitrosylated simultaneously. Specifically, the nitrosylation of the two iron sites prior to their reductive coupling to produce N2 O is cooperative. This result suggests that, in addition to any repair of iron centers (RIC) activity, YtfE acts as an NO-trapping scavenger to promote the NO to N2 O transformation under low NO flux, which precedes nitrosative stress.
- Department of Chemistry, National Tsing Hua University, Hsinchu, 30013, Taiwan.
Organizational Affiliation: