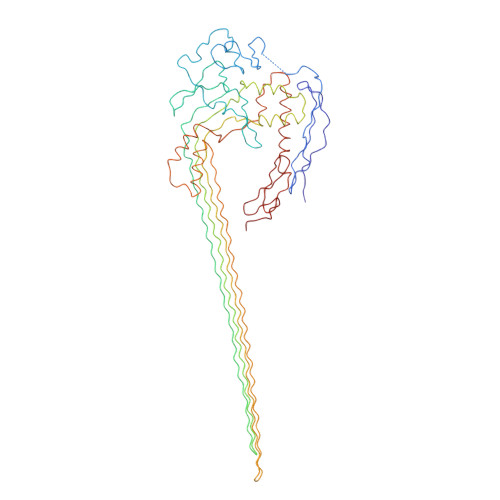Structure of the Poly-C9 Component of the Complement Membrane Attack Complex
Dudkina, N.V., Spicer, B.A., Reboul, C.F., Conroy, P.J., Lukoyanova, N., Elmlund, H., Law, R.H.P., Ekkel, S.M., Kondos, S.C., Goode, R.J.A., Ramm, G., Whisstock, J.C., Saibil, H.R., Dunstone, M.A.(2016) Nat Commun 7: 10588
- PubMed: 26841934 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms10588
- Primary Citation Related Structures:
5FMW - PubMed Abstract:
The membrane attack complex (MAC)/perforin-like protein complement component 9 (C9) is the major component of the MAC, a multi-protein complex that forms pores in the membrane of target pathogens. In contrast to homologous proteins such as perforin and the cholesterol-dependent cytolysins (CDCs), all of which require the membrane for oligomerisation, C9 assembles directly onto the nascent MAC from solution. However, the molecular mechanism of MAC assembly remains to be understood. Here we present the 8 Å cryo-EM structure of a soluble form of the poly-C9 component of the MAC. These data reveal a 22-fold symmetrical arrangement of C9 molecules that yield an 88-strand pore-forming β-barrel. The N-terminal thrombospondin-1 (TSP1) domain forms an unexpectedly extensive part of the oligomerisation interface, thus likely facilitating solution-based assembly. These TSP1 interactions may also explain how additional C9 subunits can be recruited to the growing MAC subsequent to membrane insertion.
- Department of Crystallography, Institute of Structural and Molecular Biology, Birkbeck College, London WC1E 7HX, UK.
Organizational Affiliation: