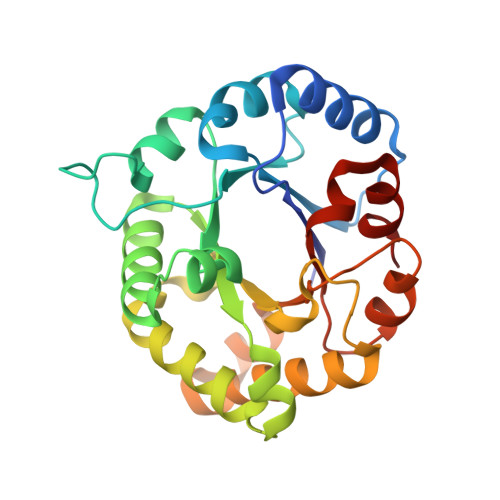
Structural insights from a novel invertebrate triosephosphate isomerase from Litopenaeus vannamei.
Lopez-Zavala, A.A., Carrasco-Miranda, J.S., Ramirez-Aguirre, C.D., Lopez-Hidalgo, M., Benitez-Cardoza, C.G., Ochoa-Leyva, A., Cardona-Felix, C.S., Diaz-Quezada, C., Rudino-Pinera, E., Sotelo-Mundo, R.R., Brieba, L.G.(2016) Biochim Biophys Acta 1864: 1696-1706
- PubMed: 27614148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbapap.2016.09.002
- Primary Citation Related Structures:
5EYW - PubMed Abstract:
Triosephosphate isomerase (TIM; EC 5.3.1.1) is a key enzyme involved in glycolysis and gluconeogenesis. Glycolysis is one of the most regulated metabolic pathways, however little is known about the structural mechanisms for its regulation in non-model organisms, like crustaceans. To understand the structure and function of this enzyme in invertebrates, we obtained the crystal structure of triosephosphate isomerase from the marine Pacific whiteleg shrimp (Litopenaeus vannamei, LvTIM) in complex with its inhibitor 2-phosphogyceric acid (2-PG) at 1.7Å resolution. LvTIM assembles as a homodimer with residues 166-176 covering the active site and residue Glu166 interacting with the inhibitor. We found that LvTIM is the least stable TIM characterized to date, with the lowest range of melting temperatures, and with the lowest activation enthalpy associated with the thermal unfolding process reported. In TIMs dimer stabilization is maintained by an interaction of loop 3 by a set of hydrophobic contacts between subunits. Within these contacts, the side chain of a hydrophobic residue of one subunit fits into a cavity created by a set of hydrophobic residues in the neighboring subunit, via a "ball and socket" interaction. LvTIM presents a Cys47 at the "ball" inter-subunit contact indicating that the character of this residue is responsible for the decrease in dimer stability. Mutational studies show that this residue plays a role in dimer stability but is not a solely determinant for dimer formation.
- Laboratorio de Estructura Biomolecular, Centro de Investigación en Alimentación y Desarrollo, A.C. (CIAD), Carretera a Ejido La Victoria Km 0.6, Apartado Postal 1735, Hermosillo, Sonora 83304, Mexico; Departamento de Ciencias Quimico Biologicas, Universidad de Sonora, Blvd. Luis Encinas y Rosales S/N, Col. Centro, Hermosillo, Sonora 83000, Mexico.
Organizational Affiliation: