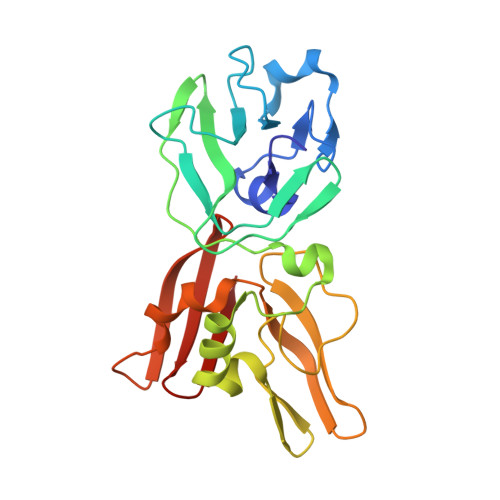
Structural Biology of the Arterivirus nsp11 Endoribonucleases.
Zhang, M., Li, X., Deng, Z., Chen, Z., Liu, Y., Gao, Y., Wu, W., Chen, Z.(2017) J Virol 91
- PubMed: 27795409 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01309-16
- Primary Citation Related Structures:
5EYI, 5HBZ, 5HC1 - PubMed Abstract:
Endoribonuclease (NendoU) is unique and conserved as a major genetic marker in nidoviruses that infect vertebrate hosts. Arterivirus nonstructural protein 11 (nsp11) was shown to have NendoU activity and play essential roles in the viral life cycle. Here, we report three crystal structures of porcine reproductive and respiratory syndrome virus (PRRSV) and equine arteritis virus (EAV) nsp11 mutants. The structures of arterivirus nsp11 contain two conserved compact domains: the N-terminal domain (NTD) and C-terminal domain (CTD). The structures of PRRSV and EAV endoribonucleases are similar and conserved in the arterivirus, but they are greatly different from that of severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS) coronaviruses (CoV), representing important human pathogens in the Nidovirales order. The catalytic center of NendoU activity is located in the CTD, where a positively charged groove is next to the key catalytic residues conserved in nidoviruses. Although the NTD is nearly identical, the catalytic region of the arterivirus nsp11 family proteins is remarkably flexible, and the oligomerization may be concentration dependent. In summary, our structures provide new insight into this key multifunctional NendoU family of proteins and lay a foundation for better understanding of the molecular mechanism and antiviral drug development. Porcine reproductive and respiratory syndrome virus (PRRSV) and equine arteritis virus are two major members of the arterivirus family. PRRSV, a leading swine pathogen, causes reproductive failure in breeding stock and respiratory tract illness in young pigs. Due to the lack of a suitable vaccine or effective drug treatment and the quick spread of these viruses, infected animals either die quickly or must be culled. PRRSV costs the swine industry around $644 million annually in the United States and almost €1.5 billion in Europe every year. To find a way to combat these viruses, we focused on the essential viral nonstructural protein 11 (nsp11). nsp11 is associated with multiple functions, such as RNA processing and suppression of the infected host innate immunity system. The three structures solved in this study provide new insight into the molecular mechanisms of this crucial protein family and will benefit the development of new treatments against these deadly viruses.
- Beijing Advanced Innovation Center for Food Nutrition and Human Health, China Agricultural University, Beijing, China.
Organizational Affiliation: