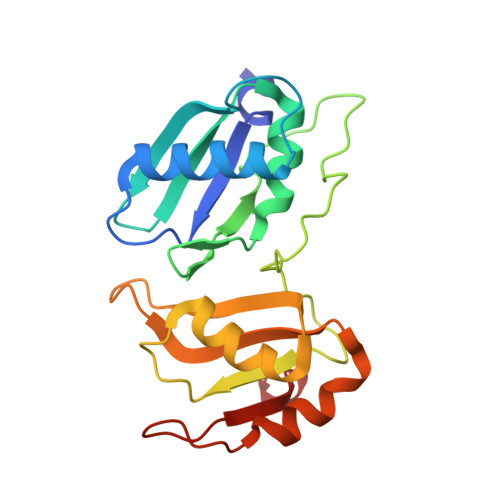
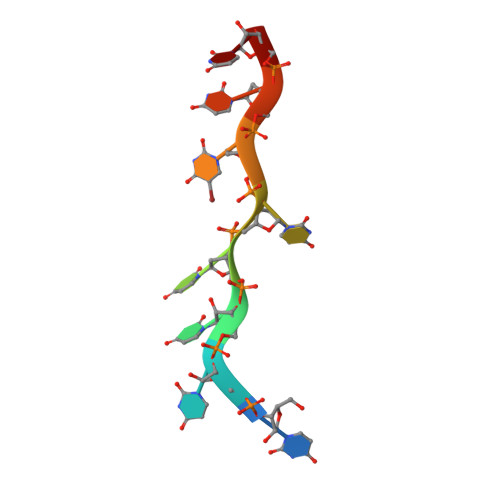
An extended U2AF(65)-RNA-binding domain recognizes the 3' splice site signal.
Agrawal, A.A., Salsi, E., Chatrikhi, R., Henderson, S., Jenkins, J.L., Green, M.R., Ermolenko, D.N., Kielkopf, C.L.(2016) Nat Commun 7: 10950-10950
- PubMed: 26952537 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms10950
- Primary Citation Related Structures:
5EV1, 5EV2, 5EV3, 5EV4 - PubMed Abstract:
How the essential pre-mRNA splicing factor U2AF(65) recognizes the polypyrimidine (Py) signals of the major class of 3' splice sites in human gene transcripts remains incompletely understood. We determined four structures of an extended U2AF(65)-RNA-binding domain bound to Py-tract oligonucleotides at resolutions between 2.0 and 1.5 Å. These structures together with RNA binding and splicing assays reveal unforeseen roles for U2AF(65) inter-domain residues in recognizing a contiguous, nine-nucleotide Py tract. The U2AF(65) linker residues between the dual RNA recognition motifs (RRMs) recognize the central nucleotide, whereas the N- and C-terminal RRM extensions recognize the 3' terminus and third nucleotide. Single-molecule FRET experiments suggest that conformational selection and induced fit of the U2AF(65) RRMs are complementary mechanisms for Py-tract association. Altogether, these results advance the mechanistic understanding of molecular recognition for a major class of splice site signals.
- Center for RNA Biology and Department of Biochemistry and Biophysics, University of Rochester School of Medicine and Dentistry, Rochester, New York 14642, USA.
Organizational Affiliation: