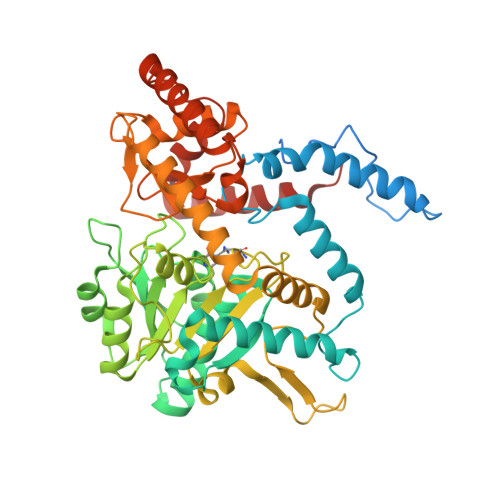
Creation of a S1P Lyase bacterial surrogate for structure-based drug design.
Argiriadi, M.A., Banach, D., Radziejewska, E., Marchie, S., DiMauro, J., Dinges, J., Dominguez, E., Hutchins, C., Judge, R.A., Queeney, K., Wallace, G., Harris, C.M.(2016) Bioorg Med Chem Lett 26: 2293-2296
- PubMed: 27013389
- DOI: https://doi.org/10.1016/j.bmcl.2016.02.084
- Primary Citation Related Structures:
5EUD, 5EUE - PubMed Abstract:
S1P Lyase (SPL) has been described as a drug target in the treatment of autoimmune diseases. It plays an important role in maintaining intracellular levels of S1P thereby affecting T cell egress from lymphoid tissues. Several groups have already published approaches to inhibit S1P Lyase with small molecules, which in turn increase endogenous S1P concentrations resulting in immunosuppression. The use of structural biology has previously aided SPL inhibitor design. Novel construct design is at times necessary to provide a reagent for protein crystallography. Here we present a chimeric bacterial protein scaffold used for protein X-ray structures in the presence of early small molecule inhibitors. Mutations were introduced to the bacterial SPL from Symbiobacterium thermophilum which mimic the human enzyme. As a result, two mutant StSPL crystal structures resolved to 2.8Å and 2.2Å resolutions were solved and provide initial structural hypotheses for an isoxazole chemical series, whose optimization is discussed in the accompanying paper.
- AbbVie Bioresearch Center, 381 Plantation Street, Worcester, MA 01605, USA. Electronic address: maria.argiriadi@abbvie.com.
Organizational Affiliation: