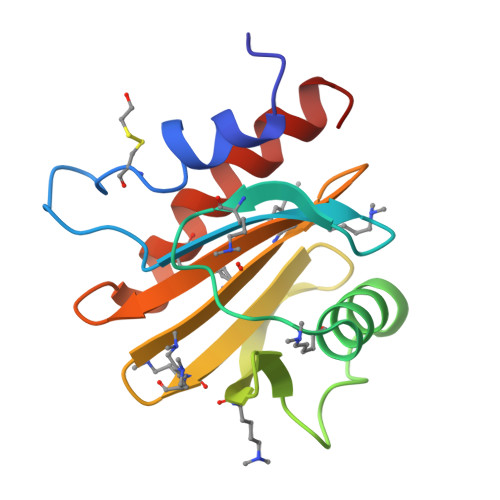
Structural, Functional, and Immunological Characterization of Profilin Panallergens Amb a 8, Art v 4, and Bet v 2.
Offermann, L.R., Schlachter, C.R., Perdue, M.L., Majorek, K.A., He, J.Z., Booth, W.T., Garrett, J., Kowal, K., Chruszcz, M.(2016) J Biological Chem 291: 15447-15459
- PubMed: 27231348 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.733659
- Primary Citation Related Structures:
5EM0, 5EM1, 5EV0, 5EVE - PubMed Abstract:
Ragweed allergens affect several million people in the United States and Canada. To date, only two ragweed allergens, Amb t 5 and Amb a 11, have their structures determined and deposited to the Protein Data Bank. Here, we present structures of methylated ragweed allergen Amb a 8, Amb a 8 in the presence of poly(l-proline), and Art v 4 (mugwort allergen). Amb a 8 and Art v 4 are panallergens belonging to the profilin family of proteins. They share significant sequence and structural similarities, which results in cross-recognition by IgE antibodies. Molecular and immunological properties of Amb a 8 and Art v 4 are compared with those of Bet v 2 (birch pollen allergen) as well as with other allergenic profilins. We purified recombinant allergens that are recognized by patient IgE and are highly cross-reactive. It was determined that the analyzed allergens are relatively unstable. Structures of Amb a 8 in complex with poly(l-proline)10 or poly(l-proline)14 are the first structures of the plant profilin in complex with proline-rich peptides. Amb a 8 binds the poly(l-proline) in a mode similar to that observed in human, mouse, and P. falciparum profilin·peptide complexes. However, only some of the residues that form the peptide binding site are conserved.
- From the Department of Chemistry and Biochemistry, University of South Carolina, Columbia, South Carolina 29208, the Department of Chemistry, Davidson College, Davidson, North Carolina 28035.
Organizational Affiliation: