CD8+T-Cell Response-Associated Evolution of Hepatitis B Virus Core Protein and Disease Progress.
Zhang, Y., Wu, Y., Deng, M., Xu, D., Li, X., Xu, Z., Hu, J., Zhang, H., Liu, K., Zhao, Y., Gao, F., Bi, S., Gao, G.F., Zhao, J., Liu, W.J., Meng, S.(2018) J Virol 92
- PubMed: 29950410 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02120-17
- Primary Citation Related Structures:
5E00, 5WSH - PubMed Abstract:
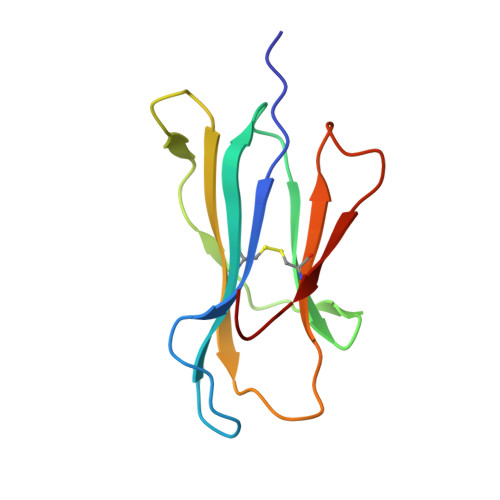
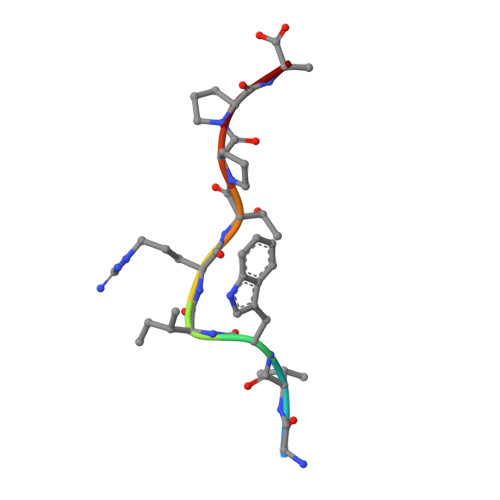
Under the immune pressure of cytotoxic T cells (CTLs), hepatitis B virus (HBV) evolves to accumulate mutations more likely within epitopes to evade immune detection. However, little is known about the specific patterns of the immune pressure-associated HBV mutation of T-cell epitopes and their link to disease progression. Here, we observed a correlation of the accumulated variants on HBV core protein (HBc) with the disease severity of HBV infection. Further analysis indicated that these substitutions were mostly located within CD8 + T-cell epitopes of HBc protein, which were systematically screened and identified in an unbiased manner in our study. From individual peptide level to the human leukocyte antigen I (HLA-I)-restricted population level, we elucidated that the mutations in these well-defined HLA-I-restricted T-cell epitopes significantly decreased antiviral activity-specific CTLs and were positively associated with clinical parameters and disease progression in HBV-infected patients. The molecular pattern for viral epitope variations based on the sequencing of 105 HBV virus genomes indicated that the C-terminal portion (Pc), especially the Pc-1 and Pc-2 positions, have the highest mutation rates. Further structural analysis of HLA-A*02 complexed to diverse CD8 + T-cell epitopes revealed that the highly variable C-terminal bulged peak of M-shaped HBc-derived epitopes are solvent exposed, and most of the CDR3βs of the T-cell receptor hover over them. These data shed light on the molecular and immunological mechanisms of T-cell immunity-associated viral evolution in hepatitis B progression, which is beneficial for designing immunotherapies and vaccines. IMPORTANCE The specific patterns of sequence polymorphisms of T-cell epitopes and the immune mechanisms of the HBV epitope mutation-linked disease progression are largely unclear. In this study, we systematically evaluated the contribution of CD8 + T cells to the disease progress-associated evolution of HBV. By evaluation of patient T-cell responses based on the peptide repertoire, we comprehensively characterized the association of clinical parameters in chronic hepatitis B with the antiviral T-cell response-associated mutations of the viruses from the single-epitope level to the overall HLA-I-restricted peptide levels. Furthermore, we investigated the molecular basis of the HLA-A2-restricted peptide immune escape and found that the solvent-exposed C-terminal portion of the epitopes is highly variable under CDR3β recognition. Our work may provide a comprehensive evaluation of viral mutations impacted by the host CTL response in HBV disease progression in the context of the full repertoire of HBc-derived epitopes.
- CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences (CAS), Beijing, China.
Organizational Affiliation: