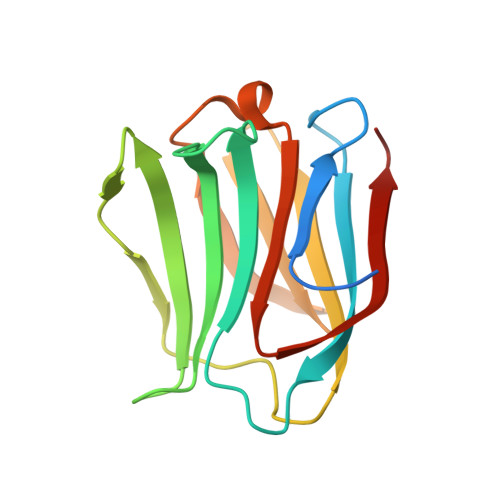
Structural characterisation of human galectin-4 N-terminal carbohydrate recognition domain in complex with glycerol, lactose, 3'-sulfo-lactose, and 2'-fucosyllactose.
Bum-Erdene, K., Leffler, H., Nilsson, U.J., Blanchard, H.(2016) Sci Rep 6: 20289-20289
- PubMed: 26828567 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep20289
- Primary Citation Related Structures:
5DUU, 5DUV, 5DUW, 5DUX - PubMed Abstract:
Galectin-4 is a tandem-repeat galectin with two distinct carbohydrate recognition domains (CRD). Galectin-4 is expressed mainly in the alimentary tract and is proposed to function as a lipid raft and adherens junction stabilizer by its glycan cross-linking capacity. Galectin-4 plays divergent roles in cancer and inflammatory conditions, either promoting or inhibiting each disease progression, depending on the specific pathological condition. The study of galectin-4's ligand-binding profile may help decipher its roles under specific conditions. Here we present the X-ray structures of human galectin-4 N-terminal CRD (galectin-4N) bound to different saccharide ligands. Galectin-4's overall fold and its core interactions to lactose are similar to other galectin CRDs. Galectin-4N recognises the sulfate cap of 3'-sulfated glycans by a weak interaction through Arg45 and two water-mediated hydrogen bonds via Trp84 and Asn49. When galectin-4N interacts with the H-antigen mimic, 2'-fucosyllactose, an interaction is formed between the ring oxygen of fucose and Arg45. The extended binding site of galectin-4N may not be well suited to the A/B-antigen determinants, α-GalNAc/α-Gal, specifically due to clashes with residue Phe47. Overall, galectin-4N favours sulfated glycans whilst galectin-4C prefers blood group determinants. However, the two CRDs of galectin-4 can, to a less extent, recognise each other's ligands.
- Institute for Glycomics, Griffith University, Gold Coast Campus, Queensland 4222, Australia.
Organizational Affiliation: