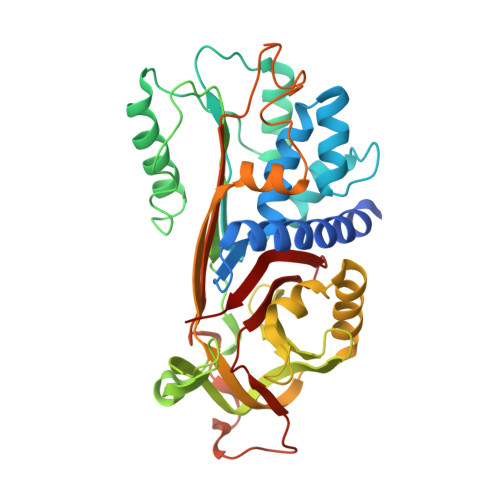
How Dextran Sulfate Affects C1-inhibitor Activity: A Model for Polysaccharide Potentiation.
Dijk, M., Holkers, J., Voskamp, P., Giannetti, B.M., Waterreus, W.J., van Veen, H.A., Pannu, N.S.(2016) Structure 24: 2182-2189
- PubMed: 27818099 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2016.09.013
- Primary Citation Related Structures:
5DU3, 5DUQ - PubMed Abstract:
C1-inhibitor is a key inhibitor of the complement and contact activation systems, and mutations in the protein can cause hereditary angioedema. Through an unknown mechanism, polysaccharides can increase C1-inhibitor activity against some of its target proteases. Here we present the crystal structures of the serine protease inhibitor (serpin) domain of active C1-inhibitor by itself and in complex with dextran sulfate. Unlike previously described interactions of serpins with polysaccharides, the structures and isothermal titration calorimetry experiments together reveal that dextran sulfate binds to C1-inhibitor's F1 helix with low affinity and does not invoke an allosteric change. Furthermore, one dextran sulfate molecule can bind multiple C1-inhibitor molecules. We propose that in a C1-inhibitor/protease/polysaccharide ternary complex, negatively charged polysaccharides link C1-inhibitor's positively charged F1 helix to positively charged autolysis loops of proteases. The proposed mechanism elegantly explains previous experiments showing that polysaccharide potentiation is increased against proteases with a greater positive charge in their autolysis loop.
- Department of Biophysical Structural Chemistry, Gorlaeus Laboratories, Leiden University, Einsteinweg 55, 2333 CC Leiden, the Netherlands.
Organizational Affiliation: