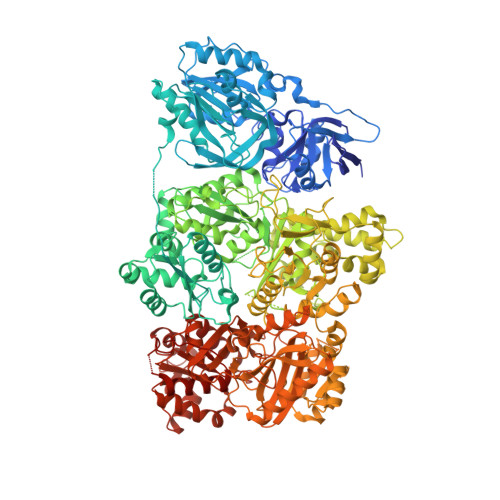
Structure of human carbamoyl phosphate synthetase: deciphering the on/off switch of human ureagenesis.
de Cima, S., Polo, L.M., Diez-Fernandez, C., Martinez, A.I., Cervera, J., Fita, I., Rubio, V.(2015) Sci Rep 5: 16950-16950
- PubMed: 26592762 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep16950
- Primary Citation Related Structures:
5DOT, 5DOU - PubMed Abstract:
Human carbamoyl phosphate synthetase (CPS1), a 1500-residue multidomain enzyme, catalyzes the first step of ammonia detoxification to urea requiring N-acetyl-L-glutamate (NAG) as essential activator to prevent ammonia/amino acids depletion. Here we present the crystal structures of CPS1 in the absence and in the presence of NAG, clarifying the on/off-switching of the urea cycle by NAG. By binding at the C-terminal domain of CPS1, NAG triggers long-range conformational changes affecting the two distant phosphorylation domains. These changes, concerted with the binding of nucleotides, result in a dramatic remodeling that stabilizes the catalytically competent conformation and the building of the ~35 Å-long tunnel that allows migration of the carbamate intermediate from its site of formation to the second phosphorylation site, where carbamoyl phosphate is produced. These structures allow rationalizing the effects of mutations found in patients with CPS1 deficiency (presenting hyperammonemia, mental retardation and even death), as exemplified here for some mutations.
- Instituto de Biomedicina de Valencia del Consejo Superior de Investigaciones Científicas (IBV-CSIC), Jaime Roig 11, 46010 Valencia, Spain.
Organizational Affiliation: