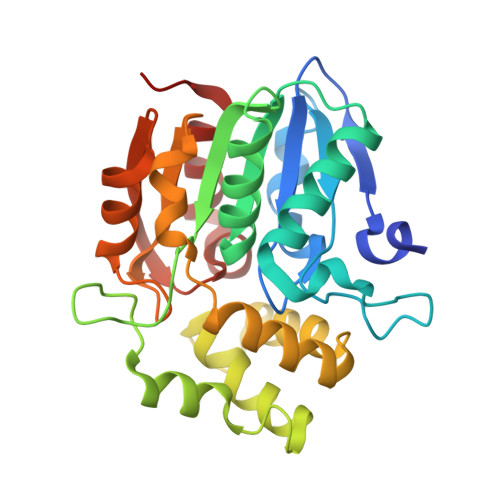
Structural basis of unique ligand specificity of KAI2-like protein from parasitic weed Striga hermonthica
Xu, Y., Miyakawa, T., Nakamura, H., Nakamura, A., Imamura, Y., Asami, T., Tanokura, M.(2016) Sci Rep 6: 31386-31386
- PubMed: 27507097 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep31386
- Primary Citation Related Structures:
5DNU, 5DNV, 5DNW - PubMed Abstract:
The perception of two plant germination inducers, karrikins and strigolactones, are mediated by the proteins KAI2 and D14. Recently, KAI2-type proteins from parasitic weeds, which are possibly related to seed germination induced by strigolactone, have been classified into three clades characterized by different responses to karrikin/strigolactone. Here we characterized a karrikin-binding protein in Striga (ShKAI2iB) that belongs to intermediate-evolving KAI2 and provided the structural bases for its karrikin-binding specificity. Binding assays showed that ShKAI2iB bound karrikins but not strigolactone, differing from other KAI2 and D14. The crystal structures of ShKAI2iB and ShKAI2iB-karrikin complex revealed obvious structural differences in a helix located at the entry of its ligand-binding cavity. This results in a smaller closed pocket, which is also the major cause of ShKAI2iB's specificity of binding karrikin. Our structural study also revealed that a few non-conserved amino acids led to the distinct ligand-binding profile of ShKAI2iB, suggesting that the evolution of KAI2 resulted in its diverse functions.
- Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: