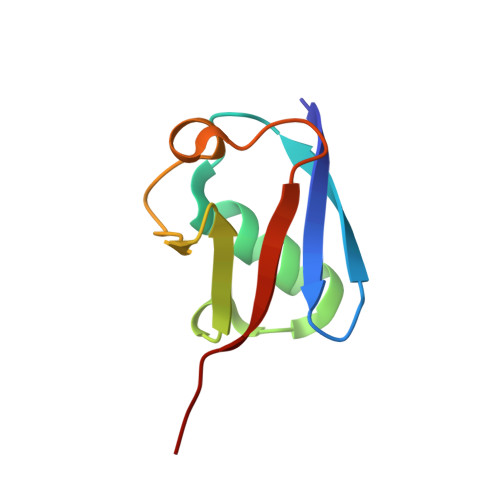
New crystal form of human ubiquitin in the presence of magnesium.
Camara-Artigas, A., Plaza-Garrido, M., Martinez-Rodriguez, S., Bacarizo, J.(2016) Acta Crystallogr F Struct Biol Commun 72: 29-35
- PubMed: 26750481 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15023390
- Primary Citation Related Structures:
5DK8 - PubMed Abstract:
Ubiquitin is a small globular protein that has a considerable number of lysine residues on its surface. This results in a high surface entropy that precludes the formation of crystal-packing interactions. To date, only a few structures of the native form of ubiquitin have been solved, and most of the crystals that led to these structures were obtained in the presence of different divalent metal cations. In this work, a new crystallographic structure of human ubiquitin solved from crystals grown in the presence of magnesium is presented. The crystals belonged to a triclinic space group, with unit-cell parameters a = 29.96, b = 30.18, c = 41.41 Å, α = 88.52, β = 79.12, γ = 67.37°. The crystal lattice is composed of stacked layers of human ubiquitin molecules with a large hydrophobic interface and a smaller polar interface in which the magnesium ion lies at the junction between adjacent layers in the crystal. The metal ion appears in a hexa-aquo coordination, which is key to facilitating the crystallization of the protein.
- Department of Chemistry and Physics, University of Almeria, Agrifood Campus of International Excellence (ceiA3), Carretera de Sacramento (s/n), 04120 Almeria, Spain.
Organizational Affiliation: