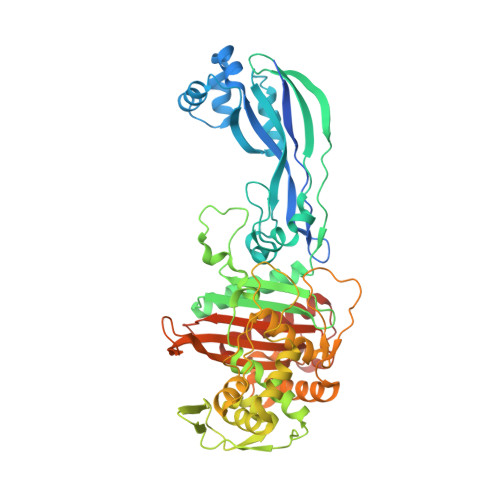
Crystal structures of penicillin-binding protein 3 in complexes with azlocillin and cefoperazone in both acylated and deacylated forms.
Ren, J., Nettleship, J.E., Males, A., Stuart, D.I., Owens, R.J.(2016) FEBS Lett 590: 288-297
- PubMed: 26823174
- DOI: https://doi.org/10.1002/1873-3468.12054
- Primary Citation Related Structures:
5DF7, 5DF8, 5DF9 - PubMed Abstract:
Penicillin-binding protein 3 (PBP3) from Pseudomonas aeruginosa is the molecular target of β-lactam-based antibiotics. Structures of PBP3 in complexes with azlocillin and cefoperazone, which are in clinical use for the treatment of pseudomonad infections, have been determined to 2.0 Å resolution. Together with data from other complexes, these structures identify a common set of residues involved in the binding of β-lactams to PBP3. Comparison of wild-type and an active site mutant (S294A) showed that increased thermal stability of PBP3 following azlocillin binding was entirely due to covalent binding to S294, whereas cefoperazone binding produces some increase in stability without the covalent link. Consistent with this, a third crystal structure was determined in which the hydrolysis product of cefoperazone was noncovalently bound in the active site of PBP3. This is the first structure of a complex between a penicillin-binding protein and cephalosporic acid and may be important in the design of new noncovalent PBP3 inhibitors.
- Division of Structural Biology, Henry Wellcome Building for Genomic Medicine, University of Oxford, Oxford, UK.
Organizational Affiliation: