A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2-ParE2 antitoxin-toxin complex.
Sterckx, Y.G., Jove, T., Shkumatov, A.V., Garcia-Pino, A., Geerts, L., De Kerpel, M., Lah, J., De Greve, H., Van Melderen, L., Loris, R.(2016) J Mol Biology 428: 1589-1603
- PubMed: 26996937 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2016.03.007
- Primary Citation Related Structures:
5CW7, 5CZE, 5CZF - PubMed Abstract:
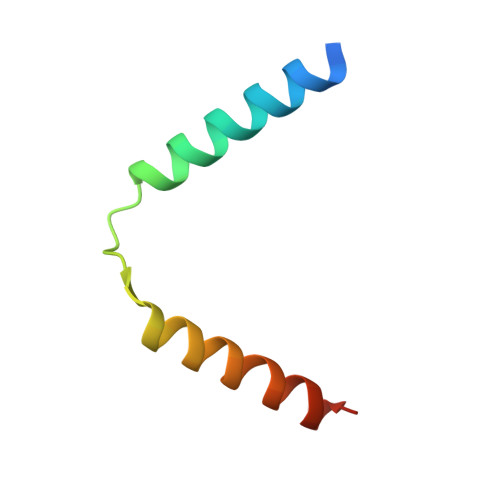
Many bacterial pathogens modulate their metabolic activity, virulence and pathogenicity through so-called "toxin-antitoxin" (TA) modules. The genome of the human pathogen Escherichia coli O157 contains two three-component TA modules related to the known parDE module. Here, we show that the toxin EcParE2 maps in a branch of the RelE/ParE toxin superfamily that is distinct from the branches that contain verified gyrase and ribosome inhibitors. The structure of EcParE2 closely resembles that of Caulobacter crescentus ParE but shows a distinct pattern of conserved surface residues, in agreement with its apparent inability to interact with GyrA. The antitoxin EcPaaA2 is characterized by two α-helices (H1 and H2) that serve as molecular recognition elements to wrap itself around EcParE2. Both EcPaaA2 H1 and H2 are required to sustain a high-affinity interaction with EcParE2 and for the inhibition of EcParE2-mediated killing in vivo. Furthermore, evidence demonstrates that EcPaaA2 H2, but not H1, determines specificity for EcParE2. The initially formed EcPaaA2-EcParE2 heterodimer then assembles into a hetero-hexadecamer, which is stable in solution and is formed in a highly cooperative manner. Together these findings provide novel data on quaternary structure, TA interactions and activity of a hitherto poorly characterized family of TA modules.
- Structural Biology Brussels, Department of Biotechnology, Vrije Universiteit Brussel (VUB), Pleinlaan 2, B-1050 Brussel, Belgium; Structural Biology Research Centre, VIB, Pleinlaan 2, B-1050 Brussel, Belgium.
Organizational Affiliation: