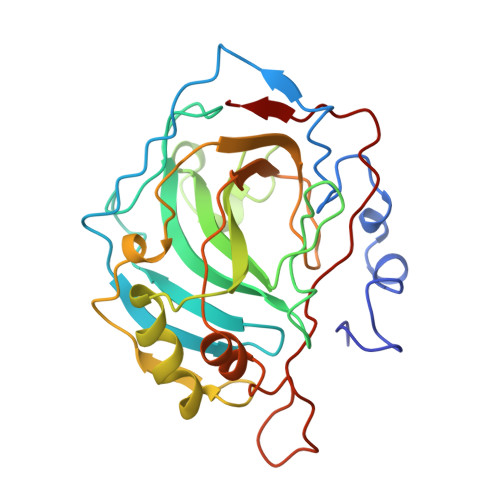
Refined structure of human carbonic anhydrase II at 2.0 A resolution.
Eriksson, A.E., Jones, T.A., Liljas, A.(1988) Proteins 4: 274-282
- PubMed: 3151019 Search on PubMed
- DOI: https://doi.org/10.1002/prot.340040406
- Primary Citation Related Structures:
1CA2, 4CAC, 5CAC - PubMed Abstract:
The structure of human erythrocytic carbonic anhydrase II has been refined by constrained and restrained structure-factor least-squares refinement at 2.0 A resolution. The conventional crystallographic R value is 17.3%. Of 167 solvent molecules associated with the protein, four are buried and stabilize secondary structure elements. The zinc ion is ligated to three histidyl residues and one water molecule in a nearly tetrahedral geometry. In addition to the zinc-bound water, seven more water molecules are identified in the active site. Assuming that Glu-106 is deprotonated at pH 8.5, some of the hydrogen bond donor-acceptor relations in the active site can be assigned and are described here in detail. The O gamma 1 atom of Thr-199 donates its proton to the O epsilon 1 atom of Glu-106 and can function as a hydrogen bond acceptor only in additional hydrogen bonds.
- Department of Molecular Biology, Biomedical Center, Uppsala, Sweden.
Organizational Affiliation: