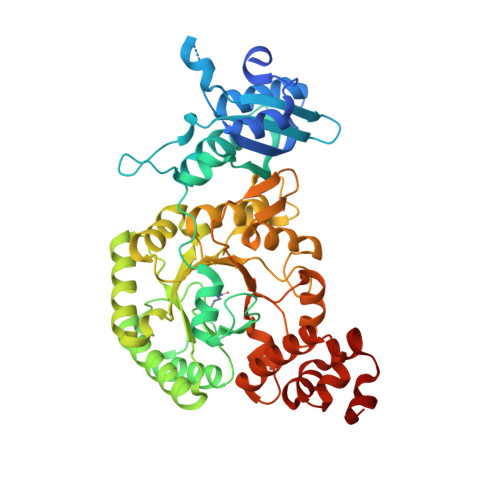
Functional metagenomic selection of ribulose 1, 5-bisphosphate carboxylase/oxygenase from uncultivated bacteria.
Varaljay, V.A., Satagopan, S., North, J.A., Witte, B., Dourado, M.N., Anantharaman, K., Arbing, M.A., McCann, S.H., Oremland, R.S., Banfield, J.F., Wrighton, K.C., Tabita, F.R.(2016) Environ Microbiol 18: 1187-1199
- PubMed: 26617072 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/1462-2920.13138
- Primary Citation Related Structures:
5C2C, 5C2G - PubMed Abstract:
Ribulose 1,5-bisphosphate carboxylase/oxygenase (RubisCO) is a critical yet severely inefficient enzyme that catalyses the fixation of virtually all of the carbon found on Earth. Here, we report a functional metagenomic selection that recovers physiologically active RubisCO molecules directly from uncultivated and largely unknown members of natural microbial communities. Selection is based on CO2 -dependent growth in a host strain capable of expressing environmental deoxyribonucleic acid (DNA), precluding the need for pure cultures or screening of recombinant clones for enzymatic activity. Seventeen functional RubisCO-encoded sequences were selected using DNA extracted from soil and river autotrophic enrichments, a photosynthetic biofilm and a subsurface groundwater aquifer. Notably, three related form II RubisCOs were recovered which share high sequence similarity with metagenomic scaffolds from uncultivated members of the Gallionellaceae family. One of the Gallionellaceae RubisCOs was purified and shown to possess CO2 /O2 specificity typical of form II enzymes. X-ray crystallography determined that this enzyme is a hexamer, only the second form II multimer ever solved and the first RubisCO structure obtained from an uncultivated bacterium. Functional metagenomic selection leverages natural biological diversity and billions of years of evolution inherent in environmental communities, providing a new window into the discovery of CO2 -fixing enzymes not previously characterized.
- Department of Microbiology, The Ohio State University, Columbus, OH, 43210, USA.
Organizational Affiliation: