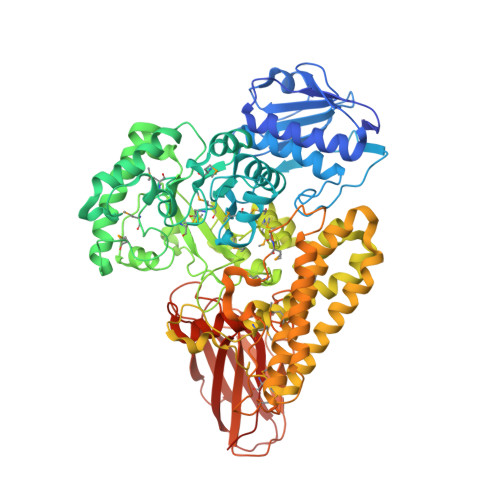
Structural and Functional Characterization of a Novel Family GH115 4-O-Methyl-alpha-Glucuronidase with Specificity for Decorated Arabinogalactans.
Aalbers, F., Turkenburg, J.P., Davies, G.J., Dijkhuizen, L., Lammerts van Bueren, A.(2015) J Mol Biology 427: 3935-3946
- PubMed: 26186997 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2015.07.006
- Primary Citation Related Structures:
5BY3 - PubMed Abstract:
Glycoside hydrolases are clustered into families based on amino acid sequence similarities, and belonging to a particular family can infer biological activity of an enzyme. Family GH115 contains α-glucuronidases where several members have been shown to hydrolyze terminal α-1,2-linked glucuronic acid and 4-O-methylated glucuronic acid from the plant cell wall polysaccharide glucuronoxylan. Other GH115 enzymes show no activity on glucuronoxylan, and therefore, it has been proposed that family GH115 may be a poly-specific family. In this study, we reveal that a putative periplasmic GH115 from the human gut symbiont Bacteroides thetaiotaomicron, BtGH115A, hydrolyzes terminal 4-O-methyl-glucuronic acid residues from decorated arabinogalactan isolated from acacia tree. The three-dimensional structure of BtGH115A reveals that BtGH115A has the same domain architecture as the other structurally characterized member of this family, BoAgu115A; however the position of the C-terminal module is altered with respect to each individual enzyme. Phylogenetic analysis of GH115 amino sequences divides the family into distinct clades that may distinguish different substrate specificities. Finally, we show that BtGH115A α-glucuronidase activity is necessary for the sequential digestion of branched galactans from acacia gum by a galactan-β-1,3-galactosidase from family GH43; however, while B. thetaiotaomicron grows on larch wood arabinogalactan, the bacterium is not able to metabolize acacia gum arabinogalactan, suggesting that BtGH115A is involved in degradation of arabinogalactan fragments liberated by other microbial species in the gastrointestinal tract.
- Microbial Physiology, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, 9747AG Groningen, The Netherlands.
Organizational Affiliation: