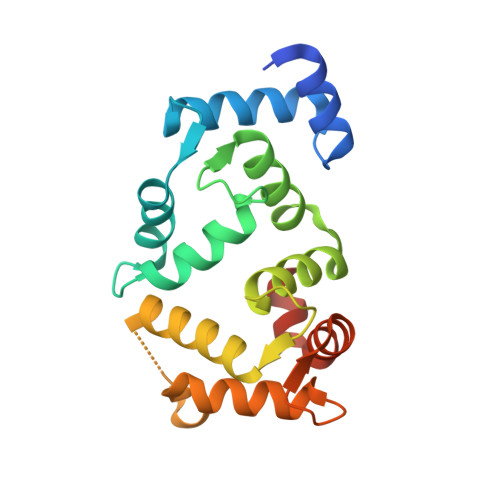
Neuronal Calcium Sensor-1 Binds the D2 Dopamine Receptor and G-Protein Coupled Receptor Kinase 1 (Grk1) Peptides Using Different Modes of Interactions.
Pandalaneni, S., Karuppiah, V., Saleem, M., Haynes, L.P., Burgoyne, R.D., Mayans, O., Derrick, J.P., Lian, L.Y.(2015) J Biological Chem 290: 18744
- PubMed: 25979333 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.627059
- Primary Citation Related Structures:
4YRU, 5AEQ, 5AER, 5AFP - PubMed Abstract:
Neuronal calcium sensor-1 (NCS-1) is the primordial member of the neuronal calcium sensor family of EF-hand Ca(2+)-binding proteins. It interacts with both the G-protein-coupled receptor (GPCR) dopamine D2 receptor (D2R), regulating its internalization and surface expression, and the cognate kinases GRK1 and GRK2. Determination of the crystal structures of Ca(2+)/NCS-1 alone and in complex with peptides derived from D2R and GRK1 reveals that the differential recognition is facilitated by the conformational flexibility of the C-lobe-binding site. We find that two copies of the D2R peptide bind within the hydrophobic crevice on Ca(2+)/NCS-1, but only one copy of the GRK1 peptide binds. The different binding modes are made possible by the C-lobe-binding site of NCS-1, which adopts alternative conformations in each complex. C-terminal residues Ser-178-Val-190 act in concert with the flexible EF3/EF4 loop region to effectively form different peptide-binding sites. In the Ca(2+)/NCS-1·D2R peptide complex, the C-terminal region adopts a 310 helix-turn-310 helix, whereas in the GRK1 peptide complex it forms an α-helix. Removal of Ser-178-Val-190 generated a C-terminal truncation mutant that formed a dimer, indicating that the NCS-1 C-terminal region prevents NCS-1 oligomerization. We propose that the flexible nature of the C-terminal region is essential to allow it to modulate its protein-binding sites and adapt its conformation to accommodate both ligands. This appears to be driven by the variability of the conformation of the C-lobe-binding site, which has ramifications for the target specificity and diversity of NCS-1.
- From the NMR Centre for Structural Biology, Institute of Integrative Biology, and.
Organizational Affiliation: