Stepping Back and Forward on Sumo Folding Evolution
Grana-Montes, R., Gallego, P., Espargaro, A., Castillo, V., Torrent, J., Lange, R., Reverter, D., Papaleo, E., Lindorff-Larsend, K., Ventura, S.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| SENTRIN-SPECIFIC PROTEASE 2 | 224 | Homo sapiens | Mutation(s): 1 EC: 3.4.22.68 (PDB Primary Data), 3.4.22 (UniProt) |  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9HC62 GTEx: ENSG00000163904 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9HC62 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| SMALL UBIQUITIN-RELATED MODIFIER 1 | 78 | Homo sapiens | Mutation(s): 2 |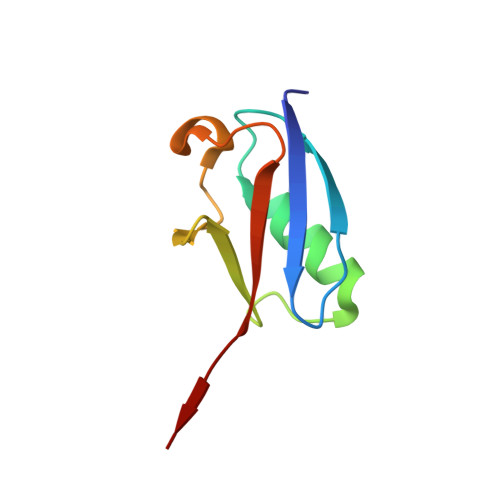  | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P63165 GTEx: ENSG00000116030 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P63165 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 113.721 | α = 90 |
| b = 119.319 | β = 89.67 |
| c = 199.84 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XPS | data reduction |
| CCP4I | data scaling |
| MOLREP | phasing |