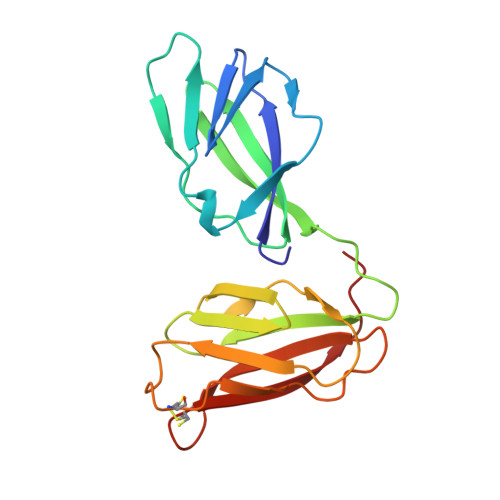
Crystal Structures of the Extracellular Domain from Pept1 and Pept2 Provide Novel Insights Into Mammalian Peptide Transport
Beale, J.H., Parker, J.L., Samsudin, F., Barrett, A.L., Senan, A., Bird, L.E., Scott, D., Owens, R.J., Sanson, M.S.P., Tucker, S.J., Meredith, D., Fowler, P.W., Newstead, S.(2015) Structure 23: 1889
- PubMed: 26320580 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.07.016
- Primary Citation Related Structures:
5A9D, 5A9H, 5A9I - PubMed Abstract:
Mammals obtain nitrogen via the uptake of di- and tri-peptides in the gastrointestinal tract through the action of PepT1 and PepT2, which are members of the POT family of proton-coupled oligopeptide transporters. PepT1 and PepT2 also play an important role in drug transport in the human body. Recent crystal structures of bacterial homologs revealed a conserved peptide-binding site and mechanism of transport. However, a key structural difference exists between bacterial and mammalian homologs with only the latter containing a large extracellular domain, the function of which is currently unknown. Here, we present the crystal structure of the extracellular domain from both PepT1 and PepT2 that reveal two immunoglobulin-like folds connected in tandem, providing structural insight into mammalian peptide transport. Functional and biophysical studies demonstrate that these domains interact with the intestinal protease trypsin, suggesting a role in clustering proteolytic activity to the site of peptide transport in eukaryotic cells.
- Department of Biochemistry, University of Oxford, Oxford OX1 3QU, UK.
Organizational Affiliation: